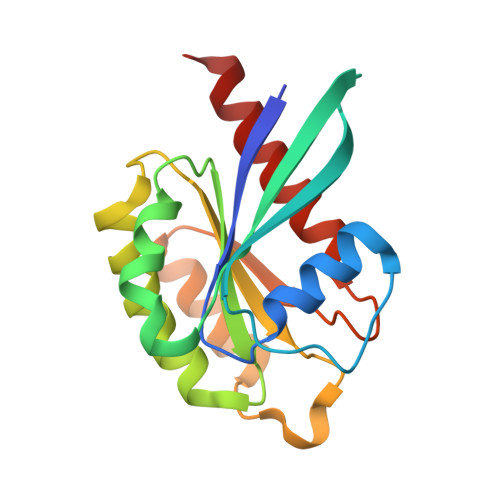
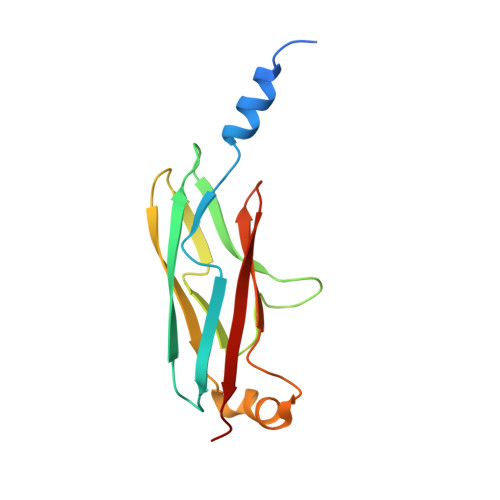
A structural basis for Lowe syndrome caused by mutations in the Rab-binding domain of OCRL1.
Hou, X., Hagemann, N., Schoebel, S., Blankenfeldt, W., Goody, R.S., Erdmann, K.S., Itzen, A.(2011) EMBO J 30: 1659-1670
- PubMed: 21378754 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2011.60
- Primary Citation Related Structures:
3QBT - PubMed Abstract:
The oculocerebrorenal syndrome of Lowe (OCRL), also called Lowe syndrome, is characterized by defects of the nervous system, the eye and the kidney. Lowe syndrome is a monogenetic X-linked disease caused by mutations of the inositol-5-phosphatase OCRL1. OCRL1 is a membrane-bound protein recruited to membranes via interaction with a variety of Rab proteins. The structural and kinetic basis of OCRL1 for the recognition of several Rab proteins is unknown. In this study, we report the crystal structure of the Rab-binding domain (RBD) of OCRL1 in complex with Rab8a and the kinetic binding analysis of OCRL1 with several Rab GTPases (Rab1b, Rab5a, Rab6a and Rab8a). In contrast to other effectors that bind their respective Rab predominantly via α-helical structure elements, the Rab-binding interface of OCRL1 consists mainly of the IgG-like β-strand structure of the ASPM-SPD-2-Hydin domain as well as one α-helix. Our results give a deeper structural understanding of disease-causing mutations of OCRL1 affecting Rab binding.
- Department of Physical Biochemistry, Max-Planck-Institute of Molecular Physiology, Dortmund, Germany.
Organizational Affiliation: