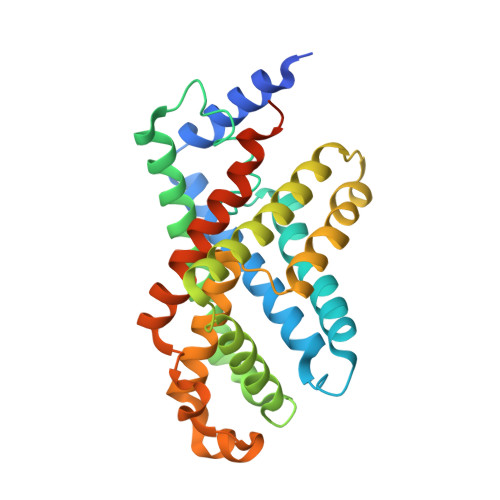
pH-dependent gating in a FocA formate channel
Lu, W., Du, J., Wacker, T., Gerbig-Smentek, E., Andrade, S.L., Einsle, O.(2011) Science 332: 352-354
- PubMed: 21493860 Search on PubMed
- DOI: https://doi.org/10.1126/science.1199098
- Primary Citation Related Structures:
3Q7K - PubMed Abstract:
The formate transporter FocA was described to switch its mode of operation from a passive export channel at high external pH to a secondary active formate/H(+) importer at low pH. The crystal structure of Salmonella typhimurium FocA at pH 4.0 shows that this switch involves a major rearrangement of the amino termini of individual protomers in the pentameric channel. The amino-terminal helices open or block transport in a concerted, cooperative action that indicates how FocA is gated in a pH-dependent way. Electrophysiological studies show that the protein acts as a specific formate channel at pH 7.0 and that it closes upon a shift of pH to 5.1.
- Lehrstuhl für Biochemie, Institut für organische Chemie und Biochemie, Albert-Ludwigs-Universität Freiburg, Albertstrasse 21, 79104 Freiburg, Germany.
Organizational Affiliation: