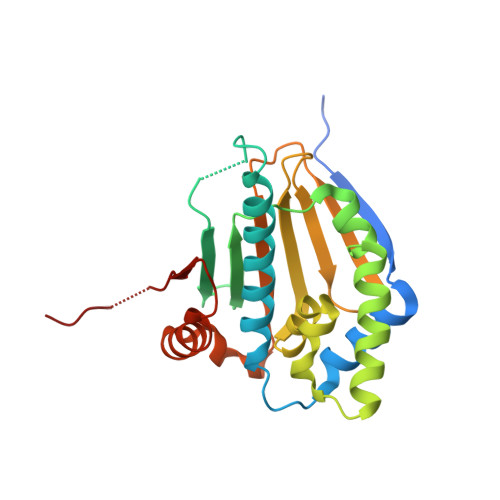
Crystal structure of the amino-terminal domain of HSP90 from Leishmania major, LMJF33.0312:M1-K213 in the presence of an inhibitor
Wernimont, A.K., Tempel, W., Lin, Y.H., Hutchinson, A., MacKenzie, F., Fairlamb, A., Cossar, D., Zhao, Y., Schapira, M., Arrowsmith, C.H., Edwards, A.M., Bountra, C., Weigelt, J., Ferguson, M.A.J., Hui, R., Pizarro, J.C., Hills, T.To be published.