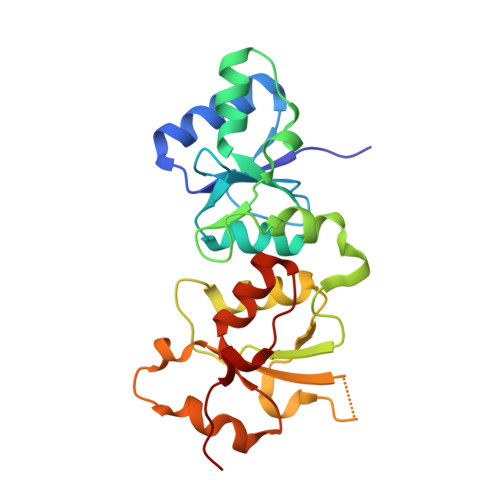
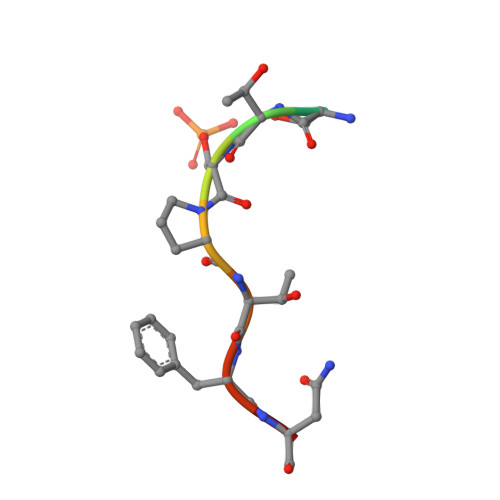
Impact of BRCA1 BRCT Domain Missense Substitutions on Phosphopeptide Recognition.
Coquelle, N., Green, R., Glover, J.N.(2011) Biochemistry 50: 4579-4589
- PubMed: 21473589 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi2003795
- Primary Citation Related Structures:
3PXA, 3PXB, 3PXC, 3PXD, 3PXE - PubMed Abstract:
The BRCA1 BRCT domain binds pSer-x-x-Phe motifs in partner proteins to regulate the cellular response to DNA damage. Approximately 120 distinct missense variants have been identified in the BRCA1 BRCT through breast cancer screening, and several of these have been linked to an increased cancer risk. Here we probe the structures and peptide-binding activities of variants that affect the BRCA1 BRCT phosphopeptide-binding groove. The results obtained from the G1656D and T1700A variants illustrate the role of Ser1655 in pSer recognition. Mutations at Arg1699 (R1699W and R1699Q) significantly reduce peptide binding through loss of contacts to the main chain of the Phe(+3) residue and, in the case of R1699W, to a destabilization of the BRCT fold. The R1835P and E1836K variants do not dramatically reduce peptide binding, in spite of the fact that these mutations significantly alter the structure of the walls of the Phe(+3) pocket.
- Department of Biochemistry, School of Medicine, University of Alberta, Edmonton, AB, Canada.
Organizational Affiliation: