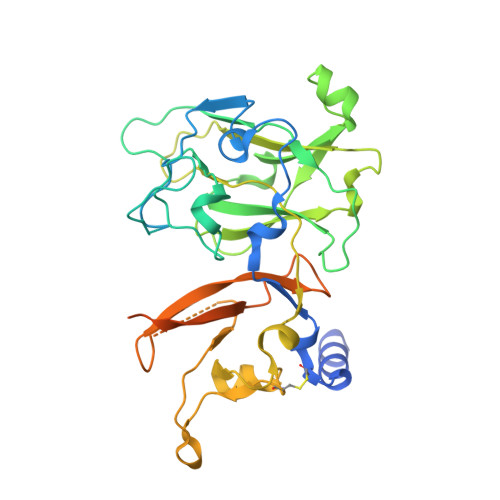
The C-Terminal Domain of the Arabinosyltransferase Mycobacterium tuberculosis EmbC Is a Lectin-Like Carbohydrate Binding Module.
Alderwick, L.J., Lloyd, G.S., Ghadbane, H., May, J.W., Bhatt, A., Eggeling, L., Futterer, K., Besra, G.S.(2011) PLoS Pathog 7: e1001299-e1001299
- PubMed: 21383969 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1001299
- Primary Citation Related Structures:
3PTY - PubMed Abstract:
The D-arabinan-containing polymers arabinogalactan (AG) and lipoarabinomannan (LAM) are essential components of the unique cell envelope of the pathogen Mycobacterium tuberculosis. Biosynthesis of AG and LAM involves a series of membrane-embedded arabinofuranosyl (Araf) transferases whose structures are largely uncharacterised, despite the fact that several of them are pharmacological targets of ethambutol, a frontline drug in tuberculosis therapy. Herein, we present the crystal structure of the C-terminal hydrophilic domain of the ethambutol-sensitive Araf transferase M. tuberculosis EmbC, which is essential for LAM synthesis. The structure of the C-terminal domain of EmbC (EmbC(CT)) encompasses two sub-domains of different folds, of which subdomain II shows distinct similarity to lectin-like carbohydrate-binding modules (CBM). Co-crystallisation with a cell wall-derived di-arabinoside acceptor analogue and structural comparison with ligand-bound CBMs suggest that EmbC(CT) contains two separate carbohydrate binding sites, associated with subdomains I and II, respectively. Single-residue substitution of conserved tryptophan residues (Trp868, Trp985) at these respective sites inhibited EmbC-catalysed extension of LAM. The same substitutions differentially abrogated binding of di- and penta-arabinofuranoside acceptor analogues to EmbC(CT), linking the loss of activity to compromised acceptor substrate binding, indicating the presence of two separate carbohydrate binding sites, and demonstrating that subdomain II indeed functions as a carbohydrate-binding module. This work provides the first step towards unravelling the structure and function of a GT-C-type glycosyltransferase that is essential in M. tuberculosis.
- School of Biosciences, University of Birmingham, Edgbaston, Birmingham, United Kingdom.
Organizational Affiliation: