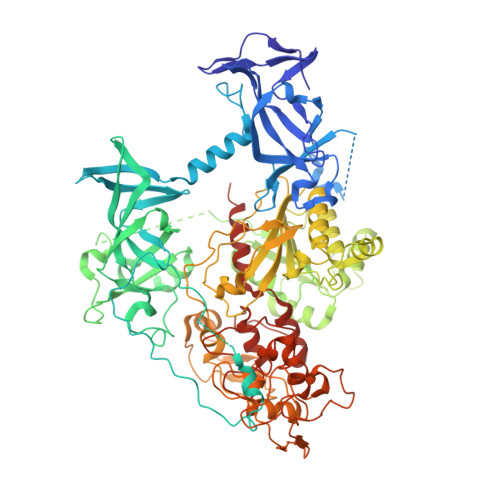
Structure of DNMT1-DNA complex reveals a role for autoinhibition in maintenance DNA methylation.
Song, J., Rechkoblit, O., Bestor, T.H., Patel, D.J.(2011) Science 331: 1036-1040
- PubMed: 21163962 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.1195380
- Primary Citation Related Structures:
3PT6, 3PT9, 3PTA - PubMed Abstract:
Maintenance of genomic methylation patterns is mediated primarily by DNA methyltransferase-1 (DNMT1). We have solved structures of mouse and human DNMT1 composed of CXXC, tandem bromo-adjacent homology (BAH1/2), and methyltransferase domains bound to DNA-containing unmethylated CpG sites. The CXXC specifically binds to unmethylated CpG dinucleotide and positions the CXXC-BAH1 linker between the DNA and the active site of DNMT1, preventing de novo methylation. In addition, a loop projecting from BAH2 interacts with the target recognition domain (TRD) of the methyltransferase, stabilizing the TRD in a retracted position and preventing it from inserting into the DNA major groove. Our studies identify an autoinhibitory mechanism, in which unmethylated CpG dinucleotides are occluded from the active site to ensure that only hemimethylated CpG dinucleotides undergo methylation.
- Structural Biology Program, Memorial Sloan-Kettering Cancer Center, New York, NY 10065, USA.
Organizational Affiliation: