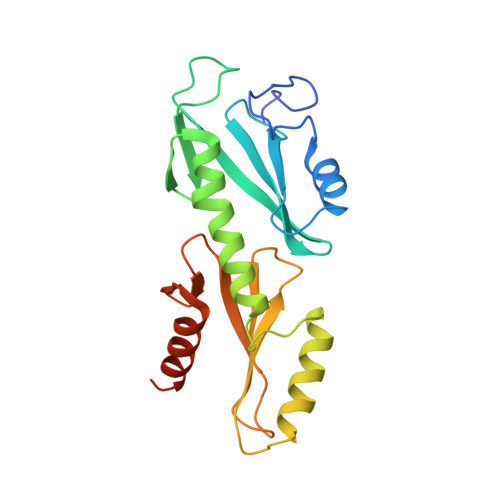
Crystal structures of the S. cerevisiae Spt6 core and C-terminal tandem SH2 domain.
Close, D., Johnson, S.J., Sdano, M.A., McDonald, S.M., Robinson, H., Formosa, T., Hill, C.P.(2011) J Mol Biology 408: 697-713
- PubMed: 21419780 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2011.03.002
- Primary Citation Related Structures:
3PSF, 3PSI, 3PSJ, 3PSK - PubMed Abstract:
The conserved and essential eukaryotic protein Spt6 functions in transcription elongation, chromatin maintenance, and RNA processing. Spt6 has three characterized functions. It is a histone chaperone capable of reassembling nucleosomes, a central component of transcription elongation complexes, and is required for recruitment of RNA processing factors to elongating RNA polymerase II (RNAPII). Here, we report multiple crystal structures of the 168-kDa Spt6 protein from Saccharomyces cerevisiae that together represent essentially all of the ordered sequence. Our two structures of the ∼900-residue core region reveal a series of putative nucleic acid and protein-protein interaction domains that fold into an elongated form that resembles the bacterial protein Tex. The similarity to a bacterial transcription factor suggests that the core domain performs nucleosome-independent activities, and as with Tex, we find that Spt6 binds DNA. Unlike Tex, however, the Spt6 S1 domain does not contribute to this activity. Crystal structures of the Spt6 C-terminal region reveal a tandem SH2 domain structure composed of two closely associated SH2 folds. One of these SH2 folds is cryptic, while the other shares striking structural similarity with metazoan SH2 domains and possesses structural features associated with the ability to bind phosphorylated substrates including phosphotyrosine. Binding studies with phosphopeptides that mimic the RNAPII C-terminal domain revealed affinities typical of other RNAPII C-terminal domain-binding proteins but did not indicate a specific interaction. Overall, these findings provide a structural foundation for understanding how Spt6 encodes several distinct functions within a single polypeptide chain.
- Department of Biochemistry, University of Utah, Salt Lake City, UT 84112-5650, USA.
Organizational Affiliation: