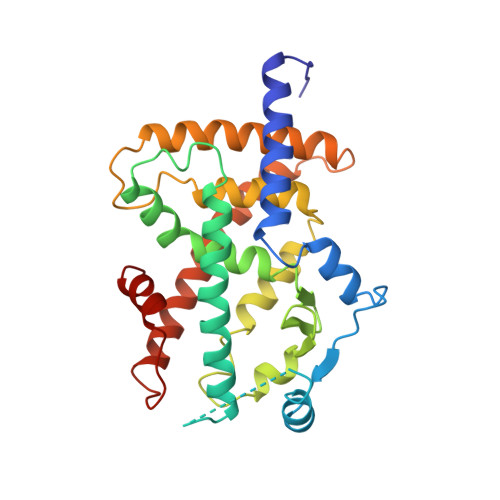
Crystal structure of the ligand binding domain of the human nuclear receptor PPARgamma.
Uppenberg, J., Svensson, C., Jaki, M., Bertilsson, G., Jendeberg, L., Berkenstam, A.(1998) J Biological Chem 273: 31108-31112
- PubMed: 9813012 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.273.47.31108
- Primary Citation Related Structures:
3PRG - PubMed Abstract:
The peroxisome proliferator-activated receptors (PPAR) are members of the nuclear receptor supergene family and are considered as key sensors of both lipid and glucose homeostasis. The role of the PPARgamma isoform in glucose metabolism is illustrated by the fact that anti-diabetic thiazolidinediones have been shown to be bona fide PPARgamma ligands. Here we report the crystal structure of apo-PPARgamma ligand binding domain (LBD) determined to 2.9-A resolution. Although the structure of apo-PPARgamma-LBD retains the overall fold described previously for other nuclear receptor LBDs, three distinct structural differences are evident. 1) The core AF-2 activation domain of apo-PPARgamma LBD is folded back toward the predicted ligand binding pocket similar to that observed in the holo-forms of other nuclear receptors. 2) The proposed ligand binding pocket of apo-PPARgamma-LBD is larger and more accessible to the surface in contrast to other LBDs. 3) The region of the LBD called the omega-loop is extended in PPARgamma and contains additional structural elements. Taken together, the apo-PPARgamma-LBD structure is in several aspects different from previously described LBDs. Given the central role of PPARgamma as a mediator in glucose regulation, the structure should be an important tool in the development of improved anti-diabetic agents.
- Department of Structural Chemistry, Pharmacia and Upjohn, Lindhagensgatan 133, S-112 87 Stockholm, Sweden. jonas.uppenberg@eu.pnu.com
Organizational Affiliation: