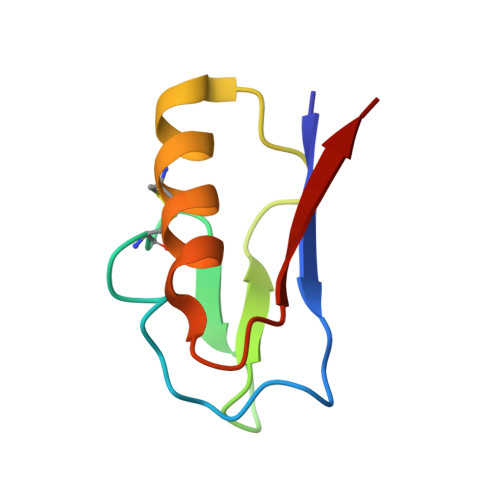
Crystal structures of the all cysteinyl coordinated D14C variant of Pyrococcus furiosus ferredoxin: [4Fe-4S] <-> [3Fe-4S] cluster conversion
Lovgreen, M.N., Martic, M., Windahl, M.S., Christensen, H.E., Harris, P.(2011) J Biol Inorg Chem 16: 763-775
- PubMed: 21484348 Search on PubMed
- DOI: https://doi.org/10.1007/s00775-011-0778-7
- Primary Citation Related Structures:
3PNI - PubMed Abstract:
The structure of the all-cysteinyl-coordinated D14C variant of [4Fe-4S] ferredoxin from the hyperthermophilic archaeon Pyrococcus furiosus has been determined to 1.7 Å resolution from a crystal belonging to space group C222(1) with two types of molecules, A and B, in the asymmetric unit. A and B molecules have different crystal packing and intramolecular disulfide bond conformation. The crystal packing reveals a β-sheet interaction between A molecules in adjacent asymmetric units, whereas B molecules are packed as monomers in a less rigid position next to the A-A extended β-sheet dimers. The A molecules contain an intramolecular disulfide bond in a double conformation with 60% occupancy left-handed and 40% occupancy right-handed spiral conformation, whereas B molecules have an intramolecular disulfide bond in a right-handed spiral conformation. The cluster in D14C [4Fe-4S] P. furiosus ferredoxin was chemically oxidized at pH 5.8 to [3Fe-4S]. For purification at pH 8.0, two forms of the protein are obtained. Mass spectrometric analysis shows that the two forms are the D14C [3Fe-4S] P. furiosus ferredoxin monomer and a disulfide-bonded dimer of D14C [3Fe-4S] P. furiosus ferredoxin. When oxidization and purification are carried out at pH 5.8, only the monomer is obtained. The crystal structure of D14C [3Fe-4S] P. furiosus ferredoxin monomer was determined to 2.8 Å resolution from a crystal belonging to space group P2(1)2(1)2(1) with two molecules in the asymmetric unit. The molecules resemble molecule A of D14C [4Fe-4S] P. furiosus ferredoxin and electron density clearly shows the presence of a [3Fe-4S] cluster.
- Department of Chemistry, Technical University of Denmark.
Organizational Affiliation: