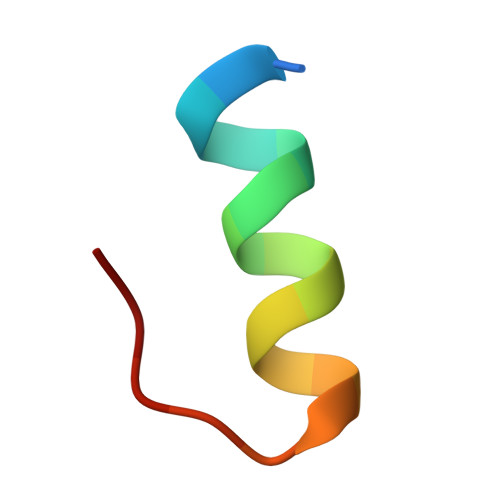
Role of the ubiquitin-like protein Hub1 in splice-site usage and alternative splicing.
Mishra, S.K., Ammon, T., Popowicz, G.M., Krajewski, M., Nagel, R.J., Ares, M., Holak, T.A., Jentsch, S.(2011) Nature 474: 173-178
- PubMed: 21614000 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature10143
- Primary Citation Related Structures:
3PLU, 3PLV - PubMed Abstract:
Alternative splicing of pre-messenger RNAs diversifies gene products in eukaryotes and is guided by factors that enable spliceosomes to recognize particular splice sites. Here we report that alternative splicing of Saccharomyces cerevisiae SRC1 pre-mRNA is promoted by the conserved ubiquitin-like protein Hub1. Structural and biochemical data show that Hub1 binds non-covalently to a conserved element termed HIND, which is present in the spliceosomal protein Snu66 in yeast and mammals, and Prp38 in plants. Hub1 binding mildly alters spliceosomal protein interactions and barely affects general splicing in S. cerevisiae. However, spliceosomes that lack Hub1, or are defective in Hub1-HIND interaction, cannot use certain non-canonical 5' splice sites and are defective in alternative SRC1 splicing. Hub1 confers alternative splicing not only when bound to HIND, but also when experimentally fused to Snu66, Prp38, or even the core splicing factor Prp8. Our study indicates a novel mechanism for splice site utilization that is guided by non-covalent modification of the spliceosome by an unconventional ubiquitin-like modifier.
- Department of Molecular Cell Biology, Max Planck Institute of Biochemistry, Am Klopferspitz 18, 82152 Martinsried, Germany.
Organizational Affiliation: