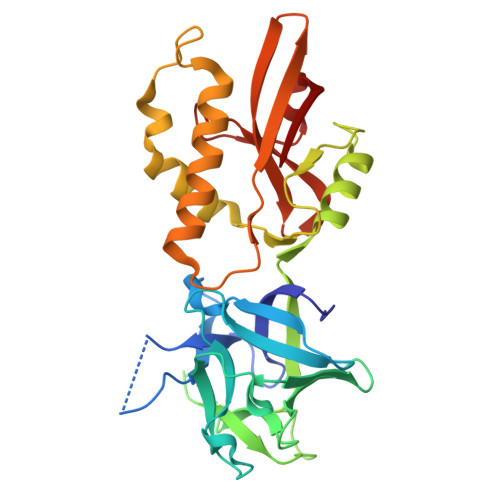
Structure and binding analysis of Polyporus squamosus lectin in complex with the Neu5Ac{alpha}2-6Gal{beta}1-4GlcNAc human-type influenza receptor.
Kadirvelraj, R., Grant, O.C., Goldstein, I.J., Winter, H.C., Tateno, H., Fadda, E., Woods, R.J.(2011) Glycobiology 21: 973-984
- PubMed: 21436237 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/glycob/cwr030
- Primary Citation Related Structures:
3PHZ - PubMed Abstract:
Glycan chains that terminate in sialic acid (Neu5Ac) are frequently the receptors targeted by pathogens for initial adhesion. Carbohydrate-binding proteins (lectins) with specificity for Neu5Ac are particularly useful in the detection and isolation of sialylated glycoconjugates, such as those associated with pathogen adhesion as well as those characteristic of several diseases including cancer. Structural studies of lectins are essential in order to understand the origin of their specificity, which is particularly important when employing such reagents as diagnostic tools. Here, we report a crystallographic and molecular dynamics (MD) analysis of a lectin from Polyporus squamosus (PSL) that is specific for glycans terminating with the sequence Neu5Acα2-6Galβ. Because of its importance as a histological reagent, the PSL structure was solved (to 1.7 Å) in complex with a trisaccharide, whose sequence (Neu5Acα2-6Galβ1-4GlcNAc) is exploited by influenza A hemagglutinin for viral adhesion to human tissue. The structural data illuminate the origin of the high specificity of PSL for the Neu5Acα2-6Gal sequence. Theoretical binding free energies derived from the MD data confirm the key interactions identified crystallographically and provide additional insight into the relative contributions from each amino acid, as well as estimates of the importance of entropic and enthalpic contributions to binding.
- Complex Carbohydrate Research Center, University of Georgia, Athens, GA 30602, USA.
Organizational Affiliation: