Molecular basis for ubiquitin and ISG15 cross-reactivity in viral ovarian tumor domains.
Akutsu, M., Ye, Y., Virdee, S., Chin, J.W., Komander, D.(2011) Proc Natl Acad Sci U S A 108: 2228-2233
- PubMed: 21266548 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1015287108
- Primary Citation Related Structures:
3PHU, 3PHW, 3PHX - PubMed Abstract:
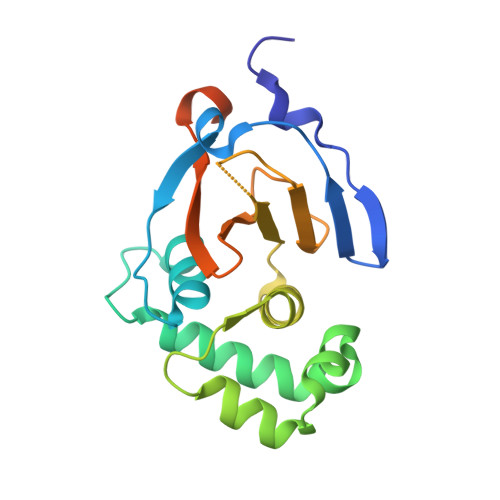
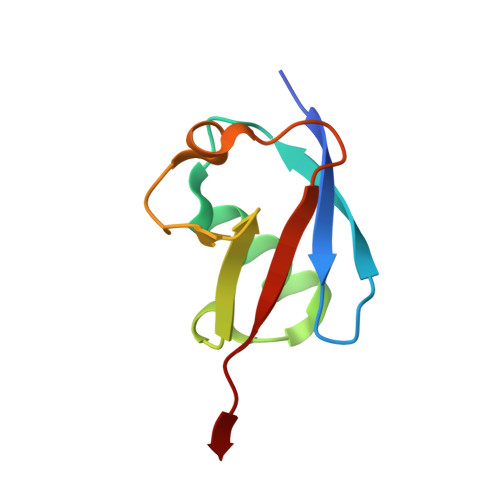
Crimean Congo hemorrhagic fever virus (CCHFV) is a deadly human pathogen that evades innate immune responses by efficiently interfering with antiviral signaling pathways mediated by NF-κB, IRF3, and IFNα/β. These pathways rely on protein ubiquitination for their activation, and one outcome is the modification of proteins with the ubiquitin (Ub)-like modifier interferon-stimulated gene (ISG)15. CCHFV and related viruses encode a deubiquitinase (DUB) of the ovarian tumor (OTU) family, which unlike eukaryotic OTU DUBs also targets ISG15 modifications. Here we characterized the viral OTU domain of CCHFV (vOTU) biochemically and structurally, revealing that it hydrolyzes four out of six tested Ub linkages, but lacks activity against linear and K29-linked Ub chains. vOTU cleaved Ub and ISG15 with similar kinetics, and we were able to understand vOTU cross-reactivity at the molecular level from crystal structures of vOTU in complex with Ub and ISG15. An N-terminal extension in vOTU not present in eukaryotic OTU binds to the hydrophobic Ile44 patch of Ub, which results in a dramatically different Ub orientation compared to a eukaryotic OTU-Ub complex. The C-terminal Ub-like fold of ISG15 (ISG15-C) adopts an equivalent binding orientation. Interestingly, ISG15-C contains an additional second hydrophobic surface that is specifically contacted by vOTU. These subtle differences in Ub/ISG15 binding allowed the design of vOTU variants specific for either Ub or ISG15, which will be useful tools to understand the relative contribution of ubiquitination vs. ISGylation in viral infection. Furthermore, the crystal structures will allow structure-based design of antiviral agents targeting this enzyme.
- Medical Research Council Laboratory of Molecular Biology, Hills Road, Cambridge CB2 0QH, United Kingdom.
Organizational Affiliation: