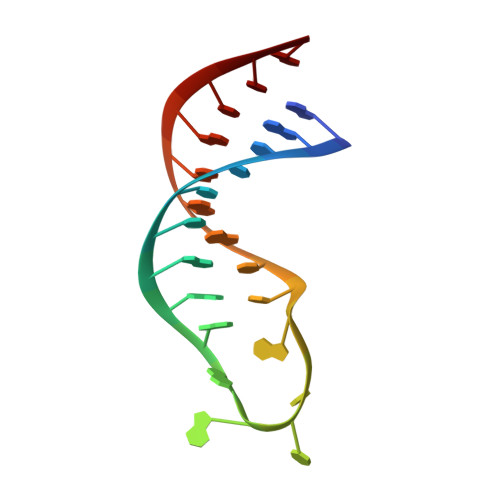
Structure of the 3'-hairpin of the TYMV pseudoknot: preformation in RNA folding.
Kolk, M.H., van der Graaf, M., Fransen, C.T., Wijmenga, S.S., Pleij, C.W., Heus, H.A., Hilbers, C.W.(1998) EMBO J 17: 7498-7504
- PubMed: 9857204
- DOI: https://doi.org/10.1093/emboj/17.24.7498
- Primary Citation Related Structures:
3PHP - PubMed Abstract:
The solution structure of an RNA-hairpin present in the pseudoknot, which is found at the 3'-terminus of turnip yellow mosaic virus genomic RNA, has been solved by nuclear magnetic resonance spectroscopy. The loop, which contains the sequence 5'-GGGUCA-3', was found to be highly structured and, contrary to expectations, does not attain its stability through GA or GC base pair formation but by triple interactions between the tilted adenosine and the minor groove sides of the first two guanosines. Interestingly, a very similar conformation was found for the cognate pseudoknot, implying that the 3'-hairpin is preformed for folding into a pseudoknotted structure. These findings suggest a mechanism of 'predetermined-fit' as a principle in RNA folding.
- NSR Center for Molecular Structure, Design and Synthesis, Laboratory of Biophysical Chemistry, University of Nijmegen, Toernooiveld, 6525 ED Nijmegen, The Netherlands.
Organizational Affiliation: