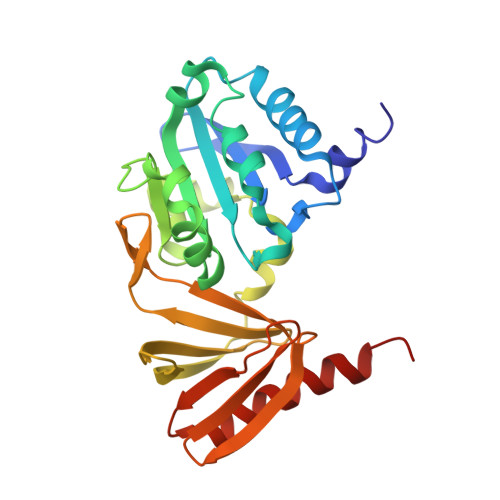
Structural and biochemical consequences of NF1 associated nontruncating mutations in the Sec14-PH module of neurofibromin.
Welti, S., Kuhn, S., D'Angelo, I., Brugger, B., Kaufmann, D., Scheffzek, K.(2011) Hum Mutat 32: 191-197
- PubMed: 21089070 Search on PubMed
- DOI: https://doi.org/10.1002/humu.21405
- Primary Citation Related Structures:
3P7Z, 3PEG, 3PG7 - PubMed Abstract:
Neurofibromatosis type 1 (NF1) is a common genetic disorder caused by alterations in the tumor suppressor gene NF1. Clinical manifestations include various neural crest derived tumors, pigmentation anomalies, bone deformations, and learning disabilities. NF1 encodes the Ras specific GTPase activating protein (RasGAP) neurofibromin, of which the central RasGAP related domain as well as a Sec14-like (residues 1560-1699) and a tightly interacting pleckstrin homology (PH)-like (1713-1818) domain are currently well defined. However, patient-derived nontruncating mutations have been reported along the whole NF1 gene, suggesting further essential protein functions. Focusing on the Sec14-PH module, we have engineered such nontruncating mutations and analyzed their implications on protein function and structure using lipid binding assays, CD spectroscopy and X-ray crystallography. Although lipid binding appears to be preserved among most nontruncating mutants, we see major structural changes for two of the alterations. Judging from these changes and our biochemical data, we suggest the presence of an intermolecular contact surface in the lid-lock region of the protein.
- European Molecular Biology Laboratory (EMBL), Structural and Computational Biology Unit, Heidelberg, Germany. welti@embl.de
Organizational Affiliation: