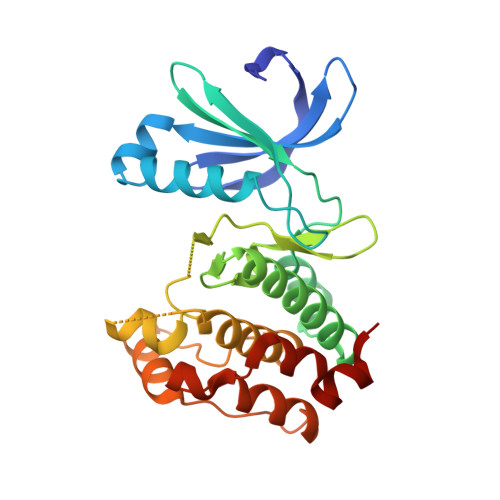
Phthalazinone pyrazoles as potent, selective, and orally bioavailable inhibitors of aurora-a kinase.
Prime, M.E., Courtney, S.M., Brookfield, F.A., Marston, R.W., Walker, V., Warne, J., Boyd, A.E., Kairies, N.A., von der Saal, W., Limberg, A., Georges, G., Engh, R.A., Goller, B., Rueger, P., Rueth, M.(2011) J Med Chem 54: 312-319
- PubMed: 21128645 Search on PubMed
- DOI: https://doi.org/10.1021/jm101346r
- Primary Citation Related Structures:
3P9J - PubMed Abstract:
The inhibition of Aurora kinases in order to arrest mitosis and subsequently inhibit tumor growth via apoptosis of proliferating cells has generated significant discussion within the literature. We report a novel class of Aurora kinase inhibitors based upon a phthalazinone pyrazole scaffold. The development of the phthalazinone template resulted in a potent Aurora-A selective series of compounds (typically >1000-fold selectivity over Aurora-B) that display good pharmacological profiles with significantly improved oral bioavailability compared to the well studied Aurora inhibitor VX-680.
- Evotec (UK) Ltd., 114 Milton Park, Abingdon, UK. Michael.Prime@evotec.com
Organizational Affiliation: