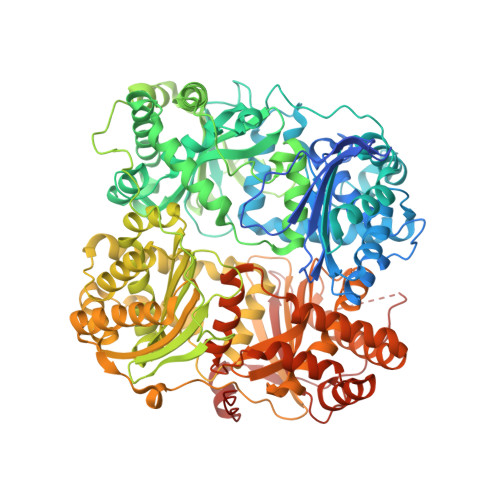
Identification of the allosteric regulatory site of insulysin.
Noinaj, N., Bhasin, S.K., Song, E.S., Scoggin, K.E., Juliano, M.A., Juliano, L., Hersh, L.B., Rodgers, D.W.(2011) PLoS One 6: e20864-e20864
- PubMed: 21731629 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0020864
- Primary Citation Related Structures:
3P7L, 3P7O - PubMed Abstract:
Insulin degrading enzyme (IDE) is responsible for the metabolism of insulin and plays a role in clearance of the Aβ peptide associated with Alzheimer's disease. Unlike most proteolytic enzymes, IDE, which consists of four structurally related domains and exists primarily as a dimer, exhibits allosteric kinetics, being activated by both small substrate peptides and polyphosphates such as ATP. The crystal structure of a catalytically compromised mutant of IDE has electron density for peptide ligands bound at the active site in domain 1 and a distal site in domain 2. Mutating residues in the distal site eliminates allosteric kinetics and activation by a small peptide, as well as greatly reducing activation by ATP, demonstrating that this site plays a key role in allostery. Comparison of the peptide bound IDE structure (using a low activity E111F IDE mutant) with unliganded wild type IDE shows a change in the interface between two halves of the clamshell-like molecule, which may enhance enzyme activity by altering the equilibrium between closed and open conformations. In addition, changes in the dimer interface suggest a basis for communication between subunits. Our findings indicate that a region remote from the active site mediates allosteric activation of insulysin by peptides. Activation may involve a small conformational change that weakens the interface between two halves of the enzyme.
- Department of Molecular and Cellular Biochemistry and Center for Structural Biology, University of Kentucky, Lexington, Kentucky, United States of America.
Organizational Affiliation: