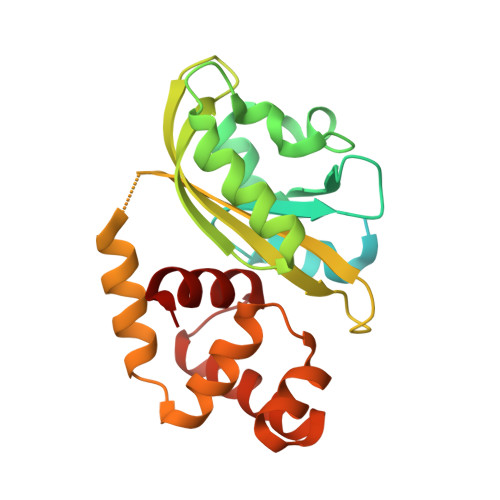
Structural basis of photosensitivity in a bacterial light-oxygen-voltage/helix-turn-helix (LOV-HTH) DNA-binding protein.
Nash, A.I., McNulty, R., Shillito, M.E., Swartz, T.E., Bogomolni, R.A., Luecke, H., Gardner, K.H.(2011) Proc Natl Acad Sci U S A 108: 9449-9454
- PubMed: 21606338 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1100262108
- Primary Citation Related Structures:
3P7N - PubMed Abstract:
Light-oxygen-voltage (LOV) domains are blue light-activated signaling modules integral to a wide range of photosensory proteins. Upon illumination, LOV domains form internal protein-flavin adducts that generate conformational changes which control effector function. Here we advance our understanding of LOV regulation with structural, biophysical, and biochemical studies of EL222, a light-regulated DNA-binding protein. The dark-state crystal structure reveals interactions between the EL222 LOV and helix-turn-helix domains that we show inhibit DNA binding. Solution biophysical data indicate that illumination breaks these interactions, freeing the LOV and helix-turn-helix domains of each other. This conformational change has a key functional effect, allowing EL222 to bind DNA in a light-dependent manner. Our data reveal a conserved signaling mechanism among diverse LOV-containing proteins, where light-induced conformational changes trigger activation via a conserved interaction surface.
- Department of Biochemistry, University of Texas Southwestern Medical Center, 5323 Harry Hines Boulevard, Dallas, TX 75390-8816, USA.
Organizational Affiliation: