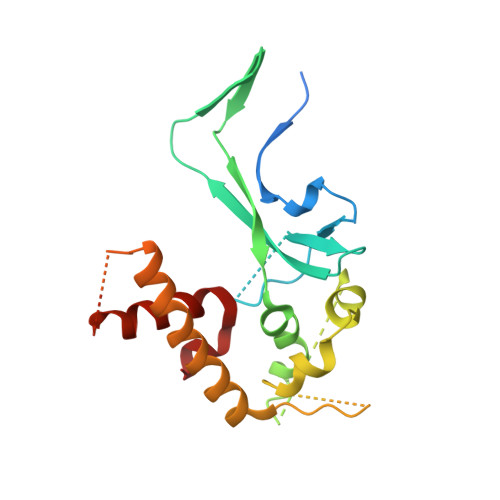
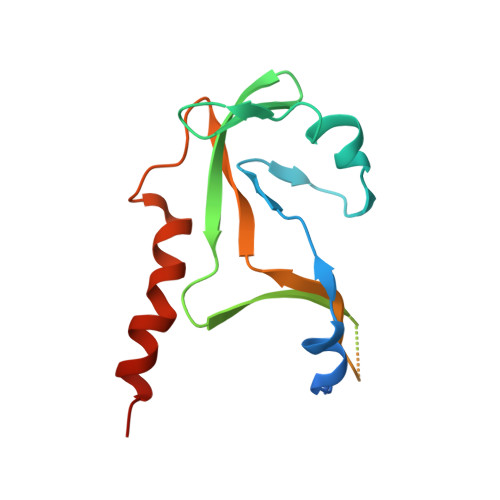
The Structure of the Human RNase H2 Complex Defines Key Interaction Interfaces Relevant to Enzyme Function and Human Disease.
Reijns, M.A., Bubeck, D., Gibson, L.C., Graham, S.C., Baillie, G.S., Jones, E.Y., Jackson, A.P.(2011) J Biological Chem 286: 10530-10539
- PubMed: 21177854 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.177394
- Primary Citation Related Structures:
3P56, 3P5J - PubMed Abstract:
Ribonuclease H2 (RNase H2) is the major nuclear enzyme involved in the degradation of RNA/DNA hybrids and removal of ribonucleotides misincorporated in genomic DNA. Mutations in each of the three RNase H2 subunits have been implicated in a human auto-inflammatory disorder, Aicardi-Goutières Syndrome (AGS). To understand how mutations impact on RNase H2 function we determined the crystal structure of the human heterotrimer. In doing so, we correct several key regions of the previously reported murine RNase H2 atomic model and provide biochemical validation for our structural model. Our results provide new insights into how the subunits are arranged to form an enzymatically active complex. In particular, we establish that the RNASEH2A C terminus is a eukaryotic adaptation for binding the two accessory subunits, with residues within it required for enzymatic activity. This C-terminal extension interacts with the RNASEH2C C terminus and both are necessary to form a stable, enzymatically active heterotrimer. Disease mutations cluster at this interface between all three subunits, destabilizing the complex and/or impairing enzyme activity. Altogether, we locate 25 out of 29 residues mutated in AGS patients, establishing a firm basis for future investigations into disease pathogenesis and function of the RNase H2 enzyme.
- Medical Research Council Human Genetics Unit, Institute of Genetics and Molecular Medicine, Western General Hospital, Edinburgh EH4 2XU, United Kingdom.
Organizational Affiliation: