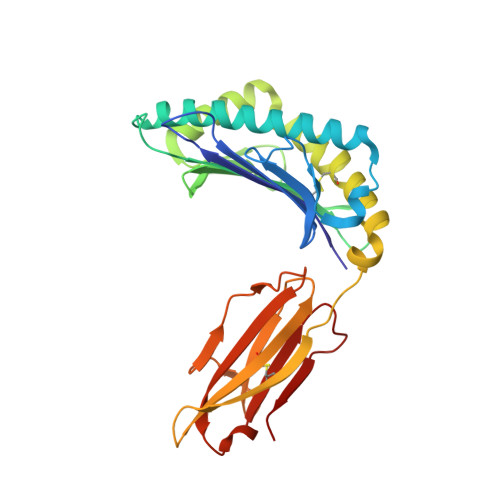
Structural insights into the binding of hepatitis B virus core peptide to HLA-A2 alleles: Towards designing better vaccines.
Liu, J., Chen, K.Y., Ren, E.C.(2011) Eur J Immunol 41: 2097-2106
- PubMed: 21538979 Search on PubMed
- DOI: https://doi.org/10.1002/eji.201041370
- Primary Citation Related Structures:
3OX8, 3OXR, 3OXS - PubMed Abstract:
Binding of specific antigenic peptides with human leukocyte antigen (HLA) molecules is a prerequisite for the initiation of T-cell responses and structural information about the peptide-HLA complex is essential for the detailed understanding of such interactions. HLA-A2 is the most prevalent HLA allele globally but aside from A*02:01 there is a significant lack of crystal structures, particularly for alleles that occur in high frequencies among Asian populations. Here, we report three HLA-A2 structures with the immunodominant hepatitis B core antigen 18-27 (HBcAg18-27) epitope, namely A*02:03, A*02:06, and A*02:07 at resolutions of 2.16, 1.70, and 1.75 Å respectively. This comparative analysis reveals that minor polymorphic residue changes between different HLA alleles can induce significant alterations in the major histocompatibility complex-peptide interface, and introduce conformational changes in the p3-p8 peptide region. Circular dichroism analysis demonstrated the HLA-A2-peptide complexes to have a hierarchy of thermostability and binding affinity in the order of A*02:06>A*02:07>A*02:01>A*02:03. Our findings provide structural insights into the varied HLA-A2 allele binding of the hepatitis B core antigen 18-27 epitope and the data suggest that chemical modifications of the peptide side chains could be a promising strategy to modulate and improve HLA-A2-peptide binding affinity for vaccine design.
- Singapore Immunology Network, A*STAR, Immunos, Singapore.
Organizational Affiliation: