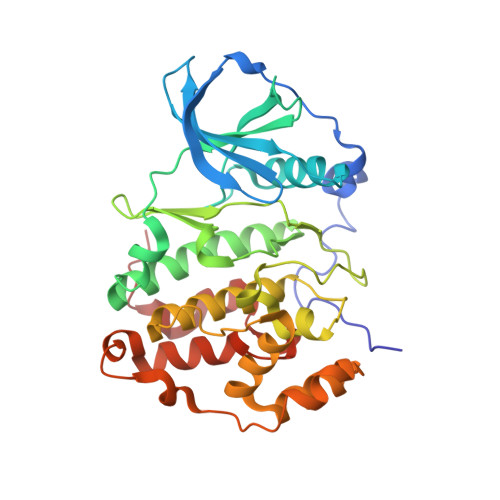
Antitumor activity of pyridocarbazole and benzopyridoindole derivatives that inhibit protein kinase CK2.
Prudent, R., Moucadel, V., Nguyen, C.H., Barette, C., Schmidt, F., Florent, J.C., Lafanechere, L., Sautel, C.F., Duchemin-Pelletier, E., Spreux, E., Filhol, O., Reiser, J.B., Cochet, C.(2010) Cancer Res 70: 9865-9874
- PubMed: 21118972 Search on PubMed
- DOI: https://doi.org/10.1158/0008-5472.CAN-10-0917
- Primary Citation Related Structures:
3OWJ, 3OWK, 3OWL - PubMed Abstract:
The alkyloid compound ellipticine derived from the berrywood tree is a topoisomerase II poison that is used in ovarian and breast cancer treatment. In this study, we report the identification of ellipticine derivatives and their tetracyclic angular benzopyridoindole analogues as novel ATP-competitive inhibitors of the protein kinase CK2. In vitro and in vivo assays showed that these compounds have a good pharmacologic profile, causing a marked inhibition of CK2 activity associated with cell cycle arrest and apoptosis in human cancer cells. Further, in vivo assays demonstrate antitumor activity in a mouse xenograft model of human glioblastoma. Finally, crystal structures of CK2-inhibitor complex provide structural insights on the molecular basis of CK2 inhibition. Our work lays the foundation for development of clinically useful CK2 inhibitors derived from a well-studied scaffold with suitable pharmacokinetics parameters.
- INSERM, U873, Grenoble, France.
Organizational Affiliation: