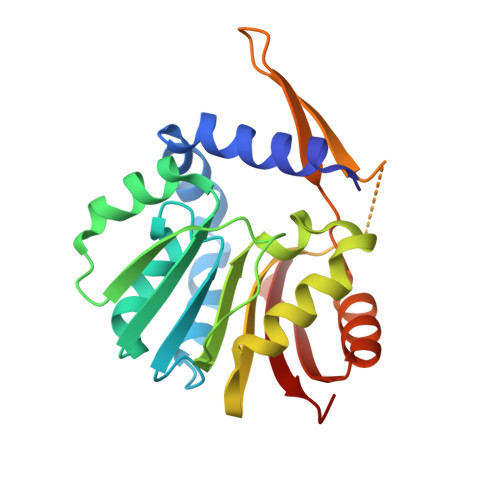
Characterization and structure of DhpI, a phosphonate O-methyltransferase involved in dehydrophos biosynthesis.
Lee, J.H., Bae, B., Kuemin, M., Circello, B.T., Metcalf, W.W., Nair, S.K., van der Donk, W.A.(2010) Proc Natl Acad Sci U S A 107: 17557-17562
- PubMed: 20876132
- DOI: https://doi.org/10.1073/pnas.1006848107
- Primary Citation of Related Structures:
3OU2, 3OU6, 3OU7 - PubMed Abstract:
Phosphonate natural products possess a range of biological activities as a consequence of their ability to mimic phosphate esters or tetrahedral intermediates formed in enzymatic reactions involved in carboxyl group metabolism. The dianionic form of these compounds at pH 7 poses a drawback with respect to their ability to mimic carboxylates and tetrahedral intermediates. Microorganisms producing phosphonates have evolved two solutions to overcome this hurdle: biosynthesis of monoanionic phosphinates containing two P-C bonds or esterification of the phosphonate group. The latter solution was first discovered for the antibiotic dehydrophos that contains a methyl ester of a phosphonodehydroalanine group. We report here the expression, purification, substrate scope, and structure of the O-methyltransferase from the dehydrophos biosynthetic gene cluster. The enzyme utilizes S-adenosylmethionine to methylate a variety of phosphonates including 1-hydroxyethylphosphonate, 1,2-dihydroxyethylphosphonate, and acetyl-1-aminoethylphosphonate. Kinetic analysis showed that the best substrates are tripeptides containing as C-terminal residue a phosphonate analog of alanine suggesting the enzyme acts late in the biosynthesis of dehydrophos. These conclusions are corroborated by the X-ray structure that reveals an active site that can accommodate a tripeptide substrate. Furthermore, the structural studies demonstrate a conformational change brought about by substrate or product binding. Interestingly, the enzyme has low substrate specificity and was used to methylate the clinical antibiotic fosfomycin and the antimalaria clinical candidate fosmidomycin, showing its promise for applications in bioengineering.
- Department of Chemistry, University of Illinois at Urbana-Champaign, 600 South Mathews Avenue, Urbana, IL 61801, USA.
Organizational Affiliation: