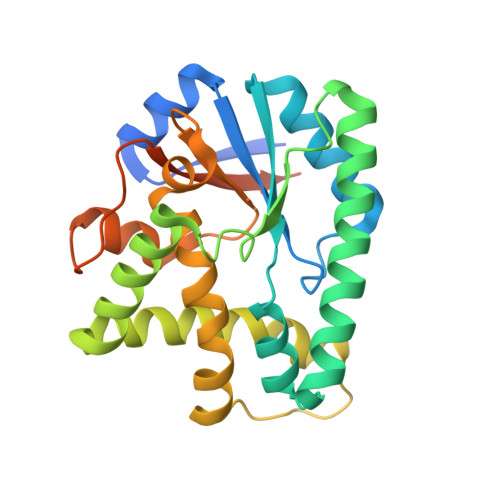
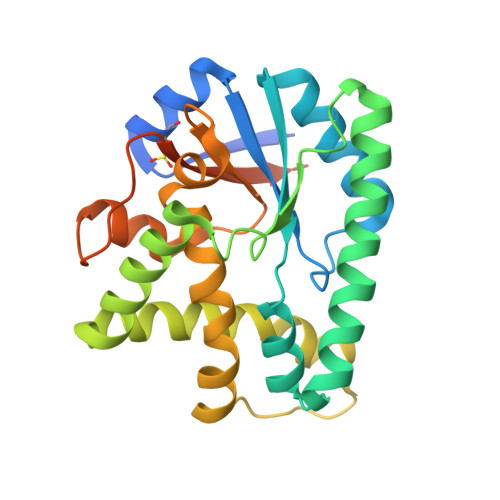
Structural basis for nonribosomal peptide synthesis by an aminoacyl-tRNA synthetase paralog.
Bonnefond, L., Arai, T., Sakaguchi, Y., Suzuki, T., Ishitani, R., Nureki, O.(2011) Proc Natl Acad Sci U S A 108: 3912-3917
- PubMed: 21325056 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1019480108
- Primary Citation Related Structures:
3OQH, 3OQI, 3OQJ, 3S7T - PubMed Abstract:
Cyclodipeptides are secondary metabolites biosynthesized by many bacteria and exhibit a wide array of biological activities. Recently, a new class of small proteins, named cyclodipeptide synthases (CDPS), which are unrelated to the typical nonribosomal peptide synthetases, was shown to generate several cyclodipeptides, using aminoacyl-tRNAs as substrates. The Mycobacterium tuberculosis CDPS, Rv2275, was found to generate cyclodityrosine through the formation of an aminoacyl-enzyme intermediate and to have a structure and oligomeric state similar to those of the class Ic aminoacyl-tRNA synthetases (aaRSs). However, the poor sequence conservation among CDPSs has raised questions about the architecture and catalytic mechanism of the identified homologs. Here we report the crystal structures of Bacillus licheniformis CDPS YvmC-Blic, in the apo form and complexed with substrate mimics, at 1.7-2.4-Å resolutions. The YvmC-Blic structure also exhibits similarity to the class Ic aaRSs catalytic domain. Our mutational analysis confirmed the importance of a set of residues for cyclodileucine formation among the conserved residues localized in the catalytic pocket. Our biochemical data indicated that YvmC-Blic binds tRNA and generates cyclodileucine as a monomer. We were also able to detect the presence of an aminoacyl-enzyme reaction intermediate, but not a dipeptide tRNA intermediate, whose existence was postulated for Rv2275. Instead, our results support a sequential catalytic mechanism for YvmC-Blic, with the successive attachment of two leucine residues on the enzyme via a conserved serine residue. Altogether, our findings suggest that all CDPS enzymes share a common aaRS-like architecture and a catalytic mechanism involving the formation of an enzyme-bound intermediate.
- Department of Biophysics and Biochemistry, Graduate School of Science, University of Tokyo, 2-11-16 Yayoi, Bunkyo-ku, 113-0032 Tokyo, Japan.
Organizational Affiliation: