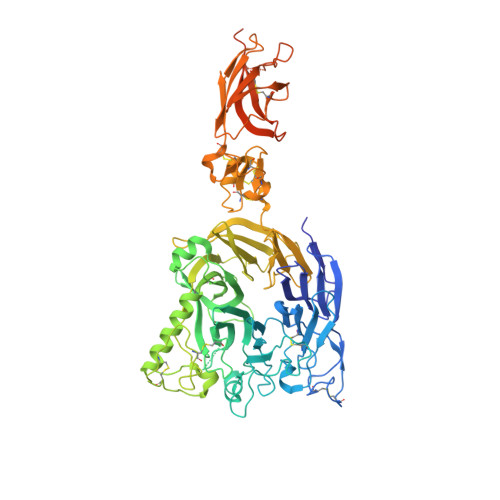
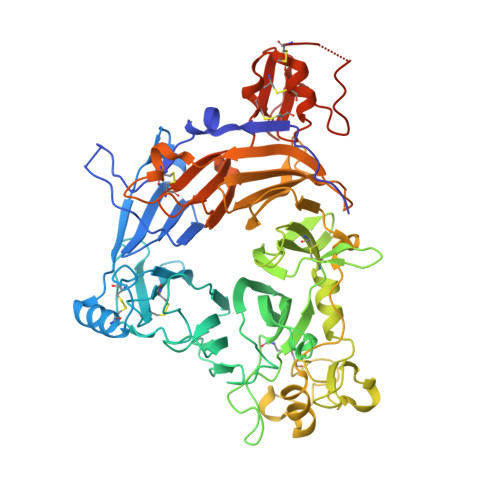
Structural basis of semaphorin-plexin signalling.
Janssen, B.J., Robinson, R.A., Perez-Branguli, F., Bell, C.H., Mitchell, K.J., Siebold, C., Jones, E.Y.(2010) Nature 467: 1118-1122
- PubMed: 20877282 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature09468
- Primary Citation Related Structures:
3OKT, 3OKW, 3OKY, 3OL2 - PubMed Abstract:
Cell-cell signalling of semaphorin ligands through interaction with plexin receptors is important for the homeostasis and morphogenesis of many tissues and is widely studied for its role in neural connectivity, cancer, cell migration and immune responses. SEMA4D and Sema6A exemplify two diverse vertebrate, membrane-spanning semaphorin classes (4 and 6) that are capable of direct signalling through members of the two largest plexin classes, B and A, respectively. In the absence of any structural information on the plexin ectodomain or its interaction with semaphorins the extracellular specificity and mechanism controlling plexin signalling has remained unresolved. Here we present crystal structures of cognate complexes of the semaphorin-binding regions of plexins B1 and A2 with semaphorin ectodomains (human PLXNB1(1-2)-SEMA4D(ecto) and murine PlxnA2(1-4)-Sema6A(ecto)), plus unliganded structures of PlxnA2(1-4) and Sema6A(ecto). These structures, together with biophysical and cellular assays of wild-type and mutant proteins, reveal that semaphorin dimers independently bind two plexin molecules and that signalling is critically dependent on the avidity of the resulting bivalent 2:2 complex (monomeric semaphorin binds plexin but fails to trigger signalling). In combination, our data favour a cell-cell signalling mechanism involving semaphorin-stabilized plexin dimerization, possibly followed by clustering, which is consistent with previous functional data. Furthermore, the shared generic architecture of the complexes, formed through conserved contacts of the amino-terminal seven-bladed β-propeller (sema) domains of both semaphorin and plexin, suggests that a common mode of interaction triggers all semaphorin-plexin based signalling, while distinct insertions within or between blades of the sema domains determine binding specificity.
- Division of Structural Biology, Wellcome Trust Centre for Human Genetics, University of Oxford, Oxford OX3 7BN, UK.
Organizational Affiliation: