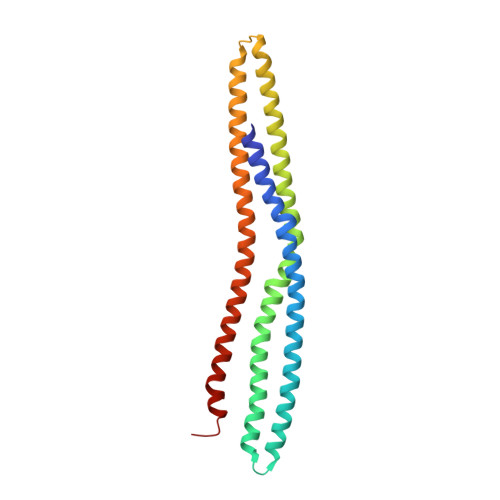
Pinkbar is an epithelial-specific BAR domain protein that generates planar membrane structures.
Pykalainen, A., Boczkowska, M., Zhao, H., Saarikangas, J., Rebowski, G., Jansen, M., Hakanen, J., Koskela, E.V., Peranen, J., Vihinen, H., Jokitalo, E., Salminen, M., Ikonen, E., Dominguez, R., Lappalainen, P.(2011) Nat Struct Mol Biol 18: 902-907
- PubMed: 21743456 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.2079
- Primary Citation Related Structures:
3OK8 - PubMed Abstract:
Bin/amphipysin/Rvs (BAR)-domain proteins sculpt cellular membranes and have key roles in processes such as endocytosis, cell motility and morphogenesis. BAR domains are divided into three subfamilies: BAR- and F-BAR-domain proteins generate positive membrane curvature and stabilize cellular invaginations, whereas I-BAR-domain proteins induce negative curvature and stabilize protrusions. We show that a previously uncharacterized member of the I-BAR subfamily, Pinkbar, is specifically expressed in intestinal epithelial cells, where it localizes to Rab13-positive vesicles and to the plasma membrane at intercellular junctions. Notably, the BAR domain of Pinkbar does not induce membrane tubulation but promotes the formation of planar membrane sheets. Structural and mutagenesis analyses reveal that the BAR domain of Pinkbar has a relatively flat lipid-binding interface and that it assembles into sheet-like oligomers in crystals and in solution, which may explain its unique membrane-deforming activity.
- Institute of Biotechnology, University of Helsinki, Helsinki, Finland.
Organizational Affiliation: