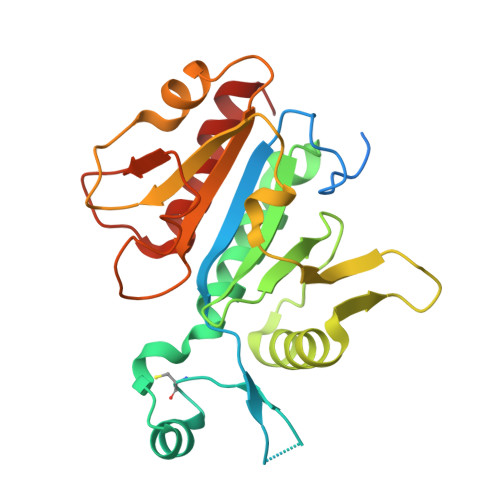
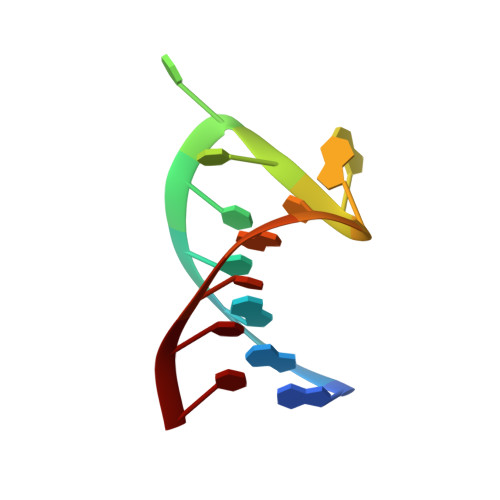
Structural insight into the functional mechanism of Nep1/Emg1 N1-specific pseudouridine methyltransferase in ribosome biogenesis.
Thomas, S.R., Keller, C.A., Szyk, A., Cannon, J.R., Laronde-Leblanc, N.A.(2011) Nucleic Acids Res 39: 2445-2457
- PubMed: 21087996 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkq1131
- Primary Citation Related Structures:
3O7B, 3OII, 3OIJ, 3OIN - PubMed Abstract:
Nucleolar Essential Protein 1 (Nep1) is required for small subunit (SSU) ribosomal RNA (rRNA) maturation and is mutated in Bowen-Conradi Syndrome. Although yeast (Saccharomyces cerevisiae) Nep1 interacts with a consensus sequence found in three regions of SSU rRNA, the molecular details of the interaction are unknown. Nep1 is a SPOUT RNA methyltransferase, and can catalyze methylation at the N1 of pseudouridine. Nep1 is also involved in assembly of Rps19, an SSU ribosomal protein. Mutations in Nep1 that result in decreased methyl donor binding do not result in lethality, suggesting that enzymatic activity may not be required for function, and RNA binding may play a more important role. To study these interactions, the crystal structures of the scNep1 dimer and its complexes with RNA were determined. The results demonstrate that Nep1 recognizes its RNA site via base-specific interactions and stabilizes a stem-loop in the bound RNA. Furthermore, the RNA structure observed contradicts the predicted structures of the Nep1-binding sites within mature rRNA, suggesting that the Nep1 changes rRNA structure upon binding. Finally, a uridine base is bound in the active site of Nep1, positioned for a methyltransfer at the C5 position, supporting its role as an N1-specific pseudouridine methyltransferase.
- Department of Chemistry and Biochemistry, Center for Biomolecular Structure and Organization, University of Maryland, College Park, MD 20742, USA.
Organizational Affiliation: