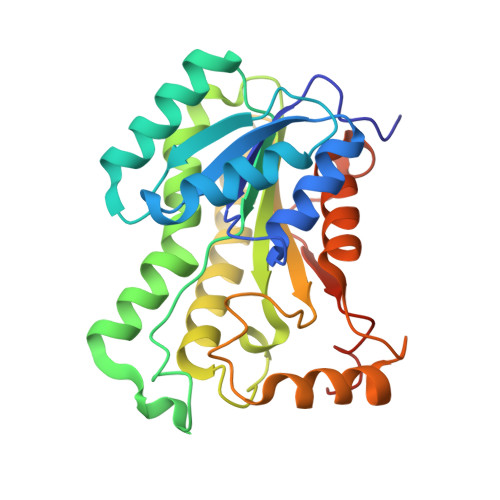
Crystal Structures of Enoyl-ACP Reductases I (FabI) and III (FabL) from B. subtilis
Kim, K.-H., Ha, B.H., Kim, S.J., Hong, S.K., Hwang, K.Y., Kim, E.E.(2011) J Mol Biology 406: 403-415
- PubMed: 21185310 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2010.12.003
- Primary Citation Related Structures:
3OIC, 3OID, 3OIF, 3OIG - PubMed Abstract:
Enoyl-[acyl carrier protein] (ACP) reductase (ENR) is a key enzyme in type II fatty acid synthesis that catalyzes the last step in each elongation cycle. Therefore, it has been considered as a target for antibiotics. However, recent studies indicate that some pathogens have more than one ENR; in particular, Bacillus subtilis has two ENRs, FabI and FabL. The crystal structures of the ternary complexes of BsFaBI and BsFabL are found as a homotetramer showing the same overall structure despite a sequence identity of only 24%. The positions of the catalytic dyad of Tyr-(Xaa)(6)-Lys in FabL are almost identical to that of FabI, but a detailed structural analysis shows that FabL shares more structural similarities with FabG and other members of the SDR (short-chain alcohol dehydrogenase/reductase) family. The apo FabL structure shows significantly different conformations at the cofactor and the substrate-binding regions, and this resulted in a totally different tetrameric arrangement reflecting the flexibility of these regions in the absence of the cofactor and substrate/inhibitor.
- Life Sciences Division, Korea Institute of Science and Technology, 39-1 Hawolkok-dong, Sungbuk-gu, Seoul 136-791, South Korea.
Organizational Affiliation: