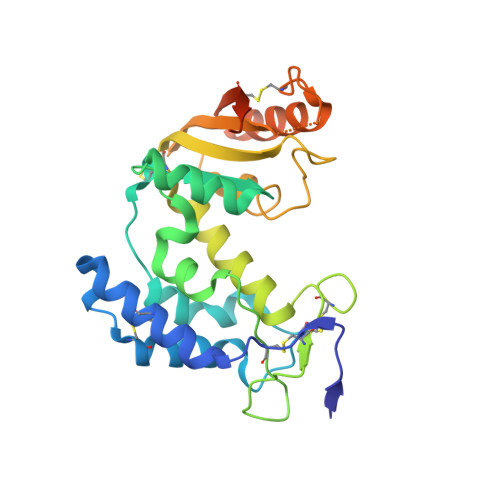
Dynamic Conformations of the CD38-Mediated NAD Cyclization Captured in a Single Crystal
Zhang, H., Graeff, R., Chen, Z., Zhang, L.R., Zhang, L.H., Lee, H.C., Hao, Q.(2011) J Mol Biology 405: 1070-1078
- PubMed: 21134381
- DOI: https://doi.org/10.1016/j.jmb.2010.11.044
- Primary Citation of Related Structures:
3OFS - PubMed Abstract:
The extracellular domain of human CD38 is a multifunctional enzyme involved in the metabolism of two Ca(2+) messengers: cyclic ADP-ribose and nicotinic acid adenine dinucleotide phosphate. When NAD is used as substrate, CD38 predominantly hydrolyzes it to ADP-ribose, with a trace amount of cyclic ADP-ribose produced through cyclization of the substrate. However, mutation of a key residue at the active site, E146, inhibits the hydrolysis activity of CD38 but greatly increases its cyclization activity. To understand the role of the residue E146 in the catalytic process, we determined the crystal structure of the E146A mutant protein with a substrate analogue, arabinosyl-2'-fluoro-deoxy-nicotinamide adenine dinucleotide. The structure captured the enzymatic reaction intermediates in six different conformations in a crystallographic asymmetric unit. The structural results indicate a folding-back process for the adenine ring of the substrate and provide the first multiple snapshots of the process. Our approach of utilizing multiple molecules in the crystallographic asymmetric unit should be generally applicable for capturing the dynamic nature of enzymatic catalysis.
- Department of Physiology, The University of Hong Kong, Hong Kong SAR, China.
Organizational Affiliation: