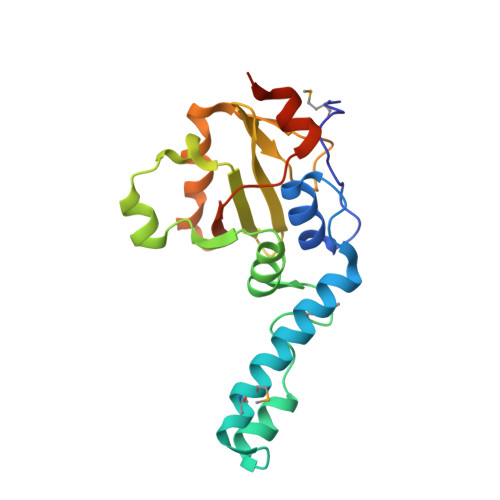
Crystal Structure of a Josephin-Ubiquitin Complex: EVOLUTIONARY RESTRAINTS ON ATAXIN-3 DEUBIQUITINATING ACTIVITY.
Weeks, S.D., Grasty, K.C., Hernandez-Cuebas, L., Loll, P.J.(2011) J Biological Chem 286: 4555-4565
- PubMed: 21118805 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.177360
- Primary Citation Related Structures:
3O65 - PubMed Abstract:
The Josephin domain is a conserved cysteine protease domain found in four human deubiquitinating enzymes: ataxin-3, the ataxin-3-like protein (ATXN3L), Josephin-1, and Josephin-2. Josephin domains from these four proteins were purified and assayed for their ability to cleave ubiquitin substrates. Reaction rates differed markedly both among the different proteins and for different substrates with a given protein. The ATXN3L Josephin domain is a significantly more efficient enzyme than the ataxin-3 domain despite their sharing 85% sequence identity. To understand the structural basis of this difference, the 2.6 Å x-ray crystal structure of the ATXN3L Josephin domain in complex with ubiquitin was determined. Although ataxin-3 and ATXN3L adopt similar folds, they bind ubiquitin in different, overlapping sites. Mutations were made in ataxin-3 at selected positions, introducing the corresponding ATXN3L residue. Only three such mutations are sufficient to increase the catalytic activity of the ataxin-3 domain to levels comparable with that of ATXN3L, suggesting that ataxin-3 has been subject to evolutionary restraints that keep its deubiquitinating activity in check.
- Department of Biochemistry and Molecular Biology, Drexel University College of Medicine, Philadelphia, Pennsylvania 19102, USA.
Organizational Affiliation: