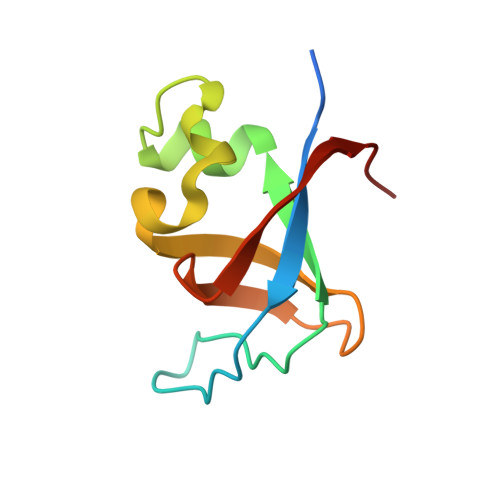
X-ray structure of the SH3 domain of the phosphoinositide 3-kinase p85 beta subunit
Chen, S., Xiao, Y., Ponnusamy, R., Tan, J., Lei, J., Hilgenfeld, R.(2011) Acta Crystallogr Sect F Struct Biol Cryst Commun 67: 1328-1333
- PubMed: 22102226 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309111031691
- Primary Citation Related Structures:
3O5Z - PubMed Abstract:
Src-homology 3 (SH3) domains are involved in extensive protein-protein interactions and constitute key elements of intracellular signal transduction. Three-dimensional structures have been reported for SH3 domains of various proteins, including the 85 kDa regulatory subunit (p85) of phosphoinositide 3-kinase. However, all of the latter structures are of p85 isoform α and no crystal structure of the SH3 domain of the equally important isoform β has been reported to date. In this structural communication, the recombinant production, crystallization and X-ray structure determination at 2.0 Å resolution of the SH3 domain of human p85β is described. The structure reveals a compact β-barrel fold very similar to that of p85α. However, binding studies with two classes of proline-rich ligand peptides demonstrate that the ligand-binding specificity differs slightly between the SH3 domains of human p85β and p85α, despite their high structural similarity.
- Institute of Biochemistry, Center for Structural and Cell Biology in Medicine, University of Lübeck, Ratzeburger Allee 160, 23538 Lübeck, Germany.
Organizational Affiliation: