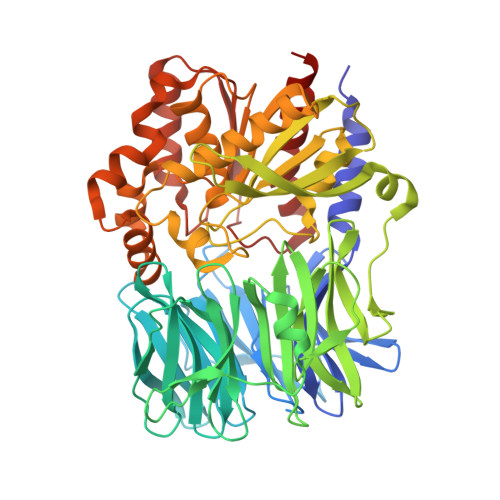
Structure and Catalysis of Acylaminoacyl Peptidase: CLOSED AND OPEN SUBUNITS OF A DIMER OLIGOPEPTIDASE.
Harmat, V., Domokos, K., Menyhard, D.K., Pallo, A., Szeltner, Z., Szamosi, I., Beke-Somfai, T., Naray-Szabo, G., Polgar, L.(2011) J Biological Chem 286: 1987-1998
- PubMed: 21084296 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.169862
- Primary Citation Related Structures:
3O4G, 3O4H, 3O4I, 3O4J - PubMed Abstract:
Acylaminoacyl peptidase from Aeropyrum pernix is a homodimer that belongs to the prolyl oligopeptidase family. The monomer subunit is composed of one hydrolase and one propeller domain. Previous crystal structure determinations revealed that the propeller domain obstructed the access of substrate to the active site of both subunits. Here we investigated the structure and the kinetics of two mutant enzymes in which the aspartic acid of the catalytic triad was changed to alanine or asparagine. Using different substrates, we have determined the pH dependence of specificity rate constants, the rate-limiting step of catalysis, and the binding of substrates and inhibitors. The catalysis considerably depended both on the kind of mutation and on the nature of the substrate. The results were interpreted in terms of alterations in the position of the catalytic histidine side chain as demonstrated with crystal structure determination of the native and two mutant structures (D524N and D524A). Unexpectedly, in the homodimeric structures, only one subunit displayed the closed form of the enzyme. The other subunit exhibited an open gate to the catalytic site, thus revealing the structural basis that controls the oligopeptidase activity. The open form of the native enzyme displayed the catalytic triad in a distorted, inactive state. The mutations affected the closed, active form of the enzyme, disrupting its catalytic triad. We concluded that the two forms are at equilibrium and the substrates bind by the conformational selection mechanism.
- Laboratory of Structural Chemistry and Biology and HAS-ELTE Protein Modeling Group, Institute of Chemistry, Eötvös Loránd University, Pázmány P. sétány 1/A, H-1117 Budapest, Hungary.
Organizational Affiliation: