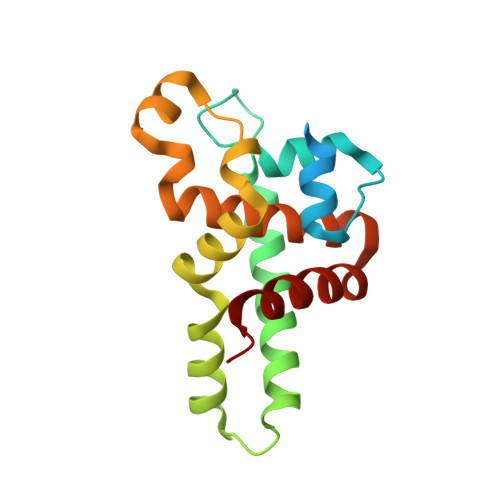
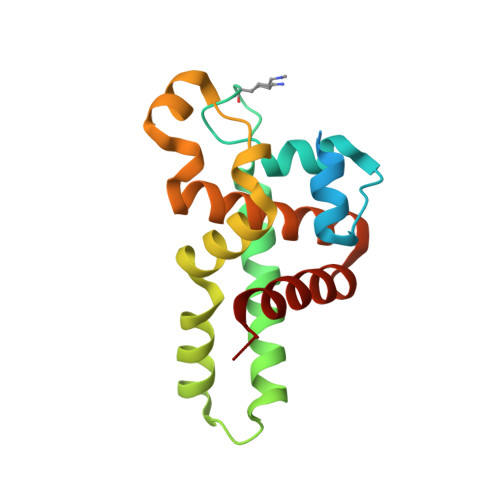
Structural Flexibility in Region Involved in Dimer Formation of Nuclease Domain of Ribonuclase III (rnc) from Campylobacter jejuni.
Minasov, G., Halavaty, A., Shuvalova, L., Dubrovska, I., Winsor, J., Papazisi, L., Anderson, W.F., Center for Structural Genomics of Infectious Diseases (CSGID)To be published.