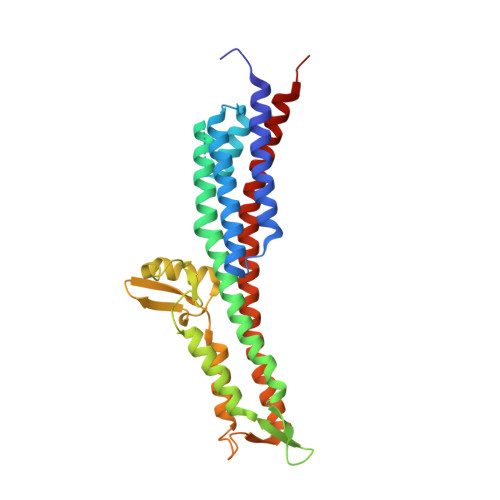
The crystal structures of the Salmonella type III secretion system tip protein SipD in complex with deoxycholate and chenodeoxycholate.
Chatterjee, S., Zhong, D., Nordhues, B.A., Battaile, K.P., Lovell, S., De Guzman, R.N.(2011) Protein Sci 20: 75-86
- PubMed: 21031487 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.537
- Primary Citation Related Structures:
3NZZ, 3O00, 3O01, 3O02 - PubMed Abstract:
The type III secretion system (T3SS) is a protein injection nanomachinery required for virulence by many human pathogenic bacteria including Salmonella and Shigella. An essential component of the T3SS is the tip protein and the Salmonella SipD and the Shigella IpaD tip proteins interact with bile salts, which serve as environmental sensors for these enteric pathogens. SipD and IpaD have long central coiled coils and their N-terminal regions form α-helical hairpins and a short helix α3 that pack against the coiled coil. Using AutoDock, others have predicted that the bile salt deoxycholate binds IpaD in a cleft formed by the α-helical hairpin and its long central coiled coil. NMR chemical shift mapping, however, indicated that the SipD residues most affected by bile salts are located in a disordered region near helix α3. Thus, how bile salts interact with SipD and IpaD is unclear. Here, we report the crystal structures of SipD in complex with the bile salts deoxycholate and chenodeoxycholate. Bile salts bind SipD in a region different from what was predicted for IpaD. In SipD, bile salts bind part of helix α3 and the C-terminus of the long central coiled coil, towards the C-terminus of the protein. We discuss the biological implication of the differences in how bile salts interact with SipD and IpaD.
- Department of Molecular Biosciences, University of Kansas, Lawrence, Kansas 66045, USA.
Organizational Affiliation: