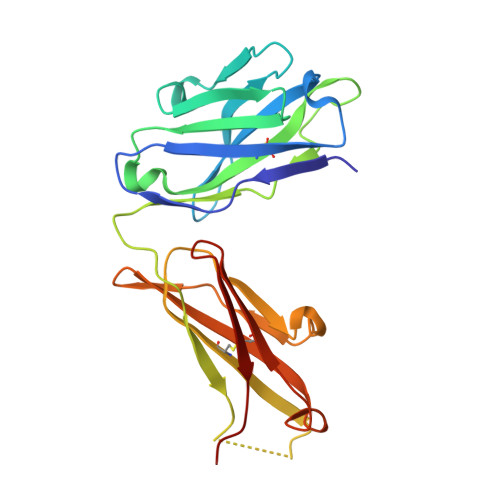
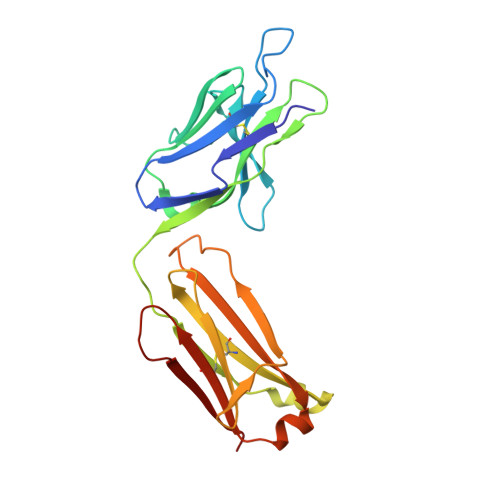
Crystal Structure of the HIV-2 Neutralizing Fab Fragment 7C8 with High Specificity to the V3 Region of gp125.
Uchtenhagen, H., Friemann, R., Raszewski, G., Spetz, A.L., Nilsson, L., Achour, A.(2011) PLoS One 6: e18767-e18767
- PubMed: 21541316 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0018767
- Primary Citation Related Structures:
3NZ8 - PubMed Abstract:
7C8 is a mouse monoclonal antibody specific for the third hypervariable region (V3) of the human immunodeficiency virus type 2 (HIV-2)-associated protein gp125. The three-dimensional crystal structure of the Fab fragment of 7C8, determined to 2.7 Å resolution, reveals a deep and narrow antigen-binding cleft with architecture appropriate for an elongated epitope. The highly hydrophobic cleft is bordered on one side by the negatively charged second complementarity determining region (CDR2) and the unusually long positively charged CDR3 of the heavy chain and, on the other side, by the CDR1 of the light chain. Analysis of 7C8 in complex with molecular models of monomeric and trimeric gp125 highlights the importance of a conserved stretch of residues FHSQ that is localized centrally on the V3 region of gp125. Furthermore, modeling also indicates that the Fab fragment neutralizes the virus by sterically impairing subsequent engagement of the gp125 trimer with the co-receptor on the target cell.
- F59 Department of Medicine, Center for Infectious Medicine, Karolinska University Hospital Huddinge, Karolinska Institutet, Stockholm, Sweden.
Organizational Affiliation: