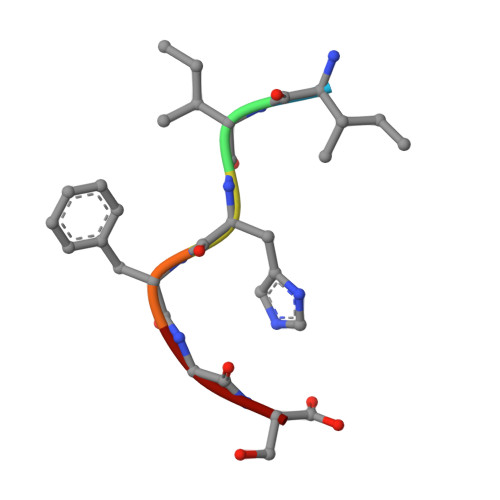
Atomic structures suggest determinants of transmission barriers in Mammalian prion disease.
Apostol, M.I., Wiltzius, J.J., Sawaya, M.R., Cascio, D., Eisenberg, D.(2011) Biochemistry 50: 2456-2463
- PubMed: 21323366 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi101803k
- Primary Citation Related Structures:
3NVE, 3NVF, 3NVG, 3NVH - PubMed Abstract:
Prion represents a unique class of pathogens devoid of nucleic acid. The deadly diseases transmitted by it between members of one species and, in certain instances, to members of other species present a public health concern. Transmissibility and the barriers to transmission between species have been suggested to arise from the degree to which a pathological protein conformation from an individual of one species can seed a pathological conformation in another species. However, this hypothesis has never been illustrated at an atomic level. Here we present three X-ray atomic structures of the same segment from human, mouse, and hamster PrP, which is critical for forming amyloid and confers species specificity in PrP seeding experiments. The structures reveal that different sequences encode different steric zippers and suggest that the degree of dissimilarity of these zipper structures gives rise to transmission barriers in prion disease, such as those that protect humans from acquiring bovine spongiform encephalopathy (BSE) and chronic wasting disease (CWD).
- Howard Hughes Medical Institute, Department of Chemistry and Biochemistry, UCLA-DOE Institute, UCLA, 611 Charles Young Drive East, Los Angeles, California 90095-1570, United States.
Organizational Affiliation: