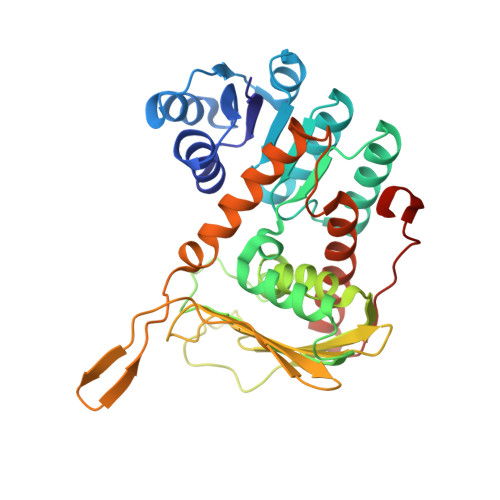
Structural investigation of myo-inositol dehydrogenase from Bacillus subtilis: implications for catalytic mechanism and inositol dehydrogenase subfamily classification.
van Straaten, K.E., Zheng, H., Palmer, D.R., Sanders, D.A.(2010) Biochem J 432: 237-247
- PubMed: 20809899 Search on PubMed
- DOI: https://doi.org/10.1042/BJ20101079
- Primary Citation Related Structures:
3MZ0, 3NT2, 3NT4, 3NT5, 3NTO, 3NTQ, 3NTR - PubMed Abstract:
Inositol dehydrogenase from Bacillus subtilis (BsIDH) is a NAD+-dependent enzyme that catalyses the oxidation of the axial hydroxy group of myo-inositol to form scyllo-inosose. We have determined the crystal structures of wild-type BsIDH and of the inactive K97V mutant in apo-, holo- and ternary complexes with inositol and inosose. BsIDH is a tetramer, with a novel arrangement consisting of two long continuous β-sheets, formed from all four monomers, in which the two central strands are crossed over to form the core of the tetramer. Each subunit in the tetramer consists of two domains: an N-terminal Rossmann fold domain containing the cofactor-binding site, and a C-terminal domain containing the inositol-binding site. Structural analysis allowed us to determine residues important in cofactor and substrate binding. Lys97, Asp172 and His176 are the catalytic triad involved in the catalytic mechanism of BsIDH, similar to what has been proposed for related enzymes and short-chain dehydrogenases. Furthermore, a conformational change in the nicotinamide ring was observed in some ternary complexes, suggesting hydride transfer to the si-face of NAD+. Finally, comparison of the structure and sequence of BsIDH with other putative inositol dehydrogenases allowed us to differentiate these enzymes into four subfamilies based on six consensus sequence motifs defining the cofactor- and substrate-binding sites.
- Department of Chemistry, University of Saskatchewan, Saskatoon, SK, Canada, S7N 5C9.
Organizational Affiliation: