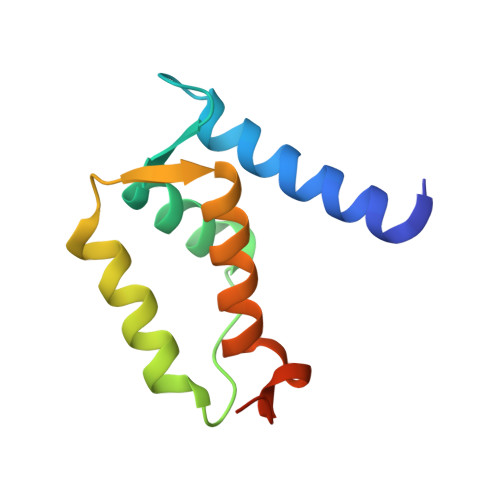
Refined crystal structures of human Ca(2+)/Zn(2+)-binding S100A3 protein characterized by two disulfide bridges
Unno, M., Kawasaki, T., Takahara, H., Heizmann, C.W., Kizawa, K.(2011) J Mol Biology 408: 477-490
- PubMed: 21377473 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2011.02.055
- Primary Citation Related Structures:
3NSI, 3NSK, 3NSL, 3NSO - PubMed Abstract:
S100A3, a member of the EF-hand-type Ca(2+)-binding S100 protein family, is unique in its exceptionally high cysteine content and Zn(2+) affinity. We produced human S100A3 protein and its mutants in insect cells using a baculovirus expression system. The purified wild-type S100A3 and the pseudo-citrullinated form (R51A) were crystallized with ammonium sulfate in N,N-bis(2-hydroxyethyl)glycine buffer and, specifically for postrefolding treatment, with Ca(2+)/Zn(2+) supplementation. We identified two previously undocumented disulfide bridges in the crystal structure of properly folded S100A3: one disulfide bridge is between Cys30 in the N-terminal pseudo-EF-hand and Cys68 in the C-terminal EF-hand (SS1), and another disulfide bridge attaches Cys99 in the C-terminal coil structure to Cys81 in helix IV (SS2). Mutational disruption of SS1 (C30A+C68A) abolished the Ca(2+) binding property of S100A3 and retarded the citrullination of Arg51 by peptidylarginine deiminase type III (PAD3), while SS2 disruption inversely increased both Ca(2+) affinity and PAD3 reactivity in vitro. Similar backbone structures of wild type, R51A, and C30A+C68A indicated that neither Arg51 conversion by PAD3 nor SS1 alters the overall dimer conformation. Comparative inspection of atomic coordinates refined to 2.15-1.40 Å resolution shows that SS1 renders the C-terminal classical Ca(2+)-binding loop flexible, which are essential for its Ca(2+) binding properties, whereas SS2 structurally shelters Arg51 in the metal-free form. We propose a model of the tetrahedral coordination of a Zn(2+) by (Cys)(3)His residues that is compatible with SS2 formation in S100A3.
- Frontier Research Center for Applied Atomic Sciences, Ibaraki University, 162-1 Shirakata, Tokai, Naka, Ibaraki 319-1106, Japan. unno19@mx.ibaraki.ac.jp
Organizational Affiliation: