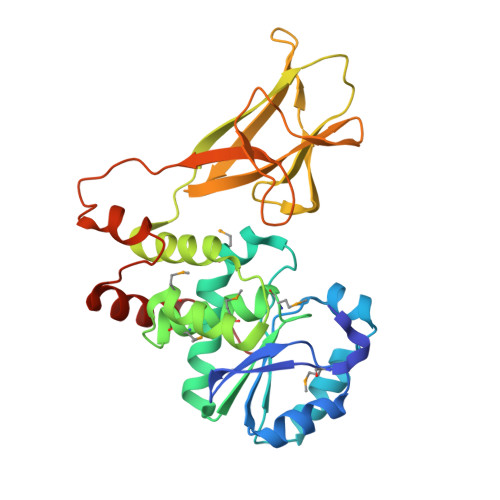
Structural basis for the glucan phosphatase activity of Starch Excess4.
Vander Kooi, C.W., Taylor, A.O., Pace, R.M., Meekins, D.A., Guo, H.F., Kim, Y., Gentry, M.S.(2010) Proc Natl Acad Sci U S A 107: 15379-15384
- PubMed: 20679247 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1009386107
- Primary Citation Related Structures:
3NME - PubMed Abstract:
Living organisms utilize carbohydrates as essential energy storage molecules. Starch is the predominant carbohydrate storage molecule in plants while glycogen is utilized in animals. Starch is a water-insoluble polymer that requires the concerted activity of kinases and phosphatases to solubilize the outer surface of the glucan and mediate starch catabolism. All known plant genomes encode the glucan phosphatase Starch Excess4 (SEX4). SEX4 can dephosphorylate both the starch granule surface and soluble phosphoglucans and is necessary for processive starch metabolism. The physical basis for the function of SEX4 as a glucan phosphatase is currently unclear. Herein, we report the crystal structure of SEX4, containing phosphatase, carbohydrate-binding, and C-terminal domains. The three domains of SEX4 fold into a compact structure with extensive interdomain interactions. The C-terminal domain of SEX4 integrally folds into the core of the phosphatase domain and is essential for its stability. The phosphatase and carbohydrate-binding domains directly interact and position the phosphatase active site toward the carbohydrate-binding site in a single continuous pocket. Mutagenesis of the phosphatase domain residue F167, which forms the base of this pocket and bridges the two domains, selectively affects the ability of SEX4 to function as a glucan phosphatase. Together, these results reveal the unique tertiary architecture of SEX4 that provides the physical basis for its function as a glucan phosphatase.
- Molecular and Cellular Biochemistry and Center for Structural Biology, University of Kentucky, Biomedical Biological Sciences Research Building, 741 South Limestone, Lexington, KY 40536-0509, USA.
Organizational Affiliation: