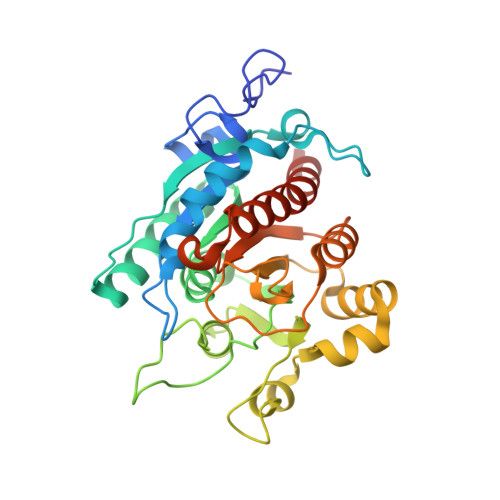
Crystal structures of Pseudomonas aeruginosa guanidinobutyrase and guanidinopropionase, members of the ureohydrolase superfamily
Lee, S.J., Kim, D.J., Kim, H.S., Lee, B.I., Yoon, H.J., Yoon, J.Y., Kim, K.H., Jang, J.Y., Im, H.N., An, D.R., Song, J.S., Kim, H.J., Suh, S.W.(2011) J Struct Biol 175: 329-338
- PubMed: 21600989 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2011.05.002
- Primary Citation Related Structures:
3NIO, 3NIP, 3NIQ - PubMed Abstract:
Pseudomonas aeruginosa guanidinobutyrase (GbuA) and guanidinopropionase (GpuA) catalyze the hydrolysis of 4-guanidinobutyrate and 3-guanidinopropionate, respectively. They belong to the ureohydrolase superfamily, which includes arginase, agmatinase, proclavaminate amidinohydrolase, and formiminoglutamase. In this study, we have determined the crystal structures of GbuA and GpuA from P. aeruginosa to provide a structural insight into their substrate specificity. Although GbuA and GpuA share a common structural fold of the typical ureohydrolase superfamily, they exhibit significant variations in two active site loops. Mutagenesis of Met161 of GbuA and Tyr157 of GpuA, both of which are located in the active site loop 1 and predicted to be involved in substrate recognition, significantly affected their enzymatic properties, implying their important roles in catalysis.
- Department of Chemistry, College of Natural Sciences, Seoul National University, Seoul 151-742, Republic of Korea.
Organizational Affiliation: