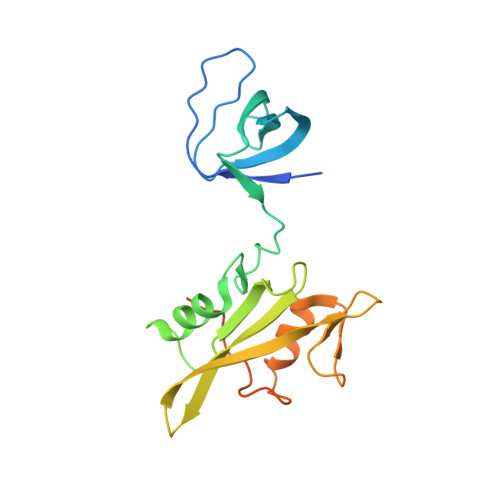
Crystal Structure of the Src Family Kinase Hck SH3-SH2 Linker Regulatory Region Supports an SH3-dominant Activation Mechanism.
Alvarado, J.J., Betts, L., Moroco, J.A., Smithgall, T.E., Yeh, J.I.(2010) J Biological Chem 285: 35455-35461
- PubMed: 20810664 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.145102
- Primary Citation Related Structures:
3NHN - PubMed Abstract:
Most mammalian cell types depend on multiple Src family kinases (SFKs) to regulate diverse signaling pathways. Strict control of SFK activity is essential for normal cellular function, and loss of kinase regulation contributes to several forms of cancer and other diseases. Previous x-ray crystal structures of the SFKs c-Src and Hck revealed that intramolecular association of their Src homology (SH) 3 domains and SH2 kinase linker regions has a key role in down-regulation of kinase activity. However, the amino acid sequence of the Hck linker represents a suboptimal ligand for the isolated SH3 domain, suggesting that it may form the polyproline type II helical conformation required for SH3 docking only in the context of the intact structure. To test this hypothesis directly, we determined the crystal structure of a truncated Hck protein consisting of the SH2 and SH3 domains plus the linker. Despite the absence of the kinase domain, the structures and relative orientations of the SH2 and SH3 domains in this shorter protein were very similar to those observed in near full-length, down-regulated Hck. However, the SH2 kinase linker adopted a modified topology and failed to engage the SH3 domain. This new structure supports the idea that these noncatalytic regions work together as a "conformational switch" that modulates kinase activity in a manner unique to the SH3 domain and linker topologies present in the intact Hck protein. Our results also provide fresh structural insight into the facile induction of Hck activity by HIV-1 Nef and other Hck SH3 domain binding proteins and implicate the existence of innate conformational states unique to individual Src family members that "fine-tune" their sensitivities to activation by SH3-based ligands.
- Department of Structural Biology, University of Pittsburgh School of Medicine, Pittsburgh, Pennsylvania 15260, USA.
Organizational Affiliation: