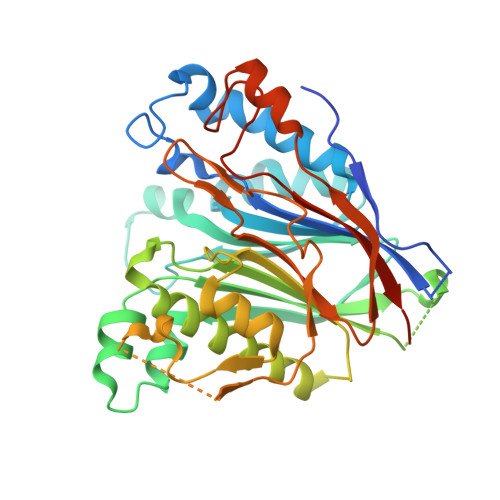
Crystal structure of the human CNOT6L nuclease domain reveals strict poly(A) substrate specificity.
Wang, H., Morita, M., Yang, X., Suzuki, T., Yang, W., Wang, J., Ito, K., Wang, Q., Zhao, C., Bartlam, M., Yamamoto, T., Rao, Z.(2010) EMBO J
- PubMed: 20628353 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2010.152
- Primary Citation Related Structures:
3NGN, 3NGO, 3NGQ - PubMed Abstract:
CCR4, an evolutionarily conserved member of the CCR4-NOT complex, is the main cytoplasmic deadenylase. It contains a C-terminal nuclease domain with homology to the endonuclease-exonuclease-phosphatase (EEP) family of enzymes. We have determined the high-resolution three-dimensional structure of the nuclease domain of CNOT6L, a human homologue of CCR4, by X-ray crystallography using the single-wavelength anomalous dispersion method. This first structure of a deadenylase belonging to the EEP family adopts a complete alpha/beta sandwich fold typical of hydrolases with highly conserved active site residues similar to APE1. The active site of CNOT6L should recognize the RNA substrate through its negatively charged surface. In vitro deadenylase assays confirm the critical active site residues and show that the nuclease domain of CNOT6L exhibits full Mg(2+)-dependent deadenylase activity with strict poly(A) RNA substrate specificity. To understand the structural basis for poly(A) RNA substrate binding, crystal structures of the CNOT6L nuclease domain have also been determined in complex with AMP and poly(A) DNA. The resulting structures suggest a molecular deadenylase mechanism involving a pentacovalent phosphate transition.
- Tianjin Key Laboratory of Protein Science, College of Life Sciences, Nankai University, Tianjin, PR China.
Organizational Affiliation: