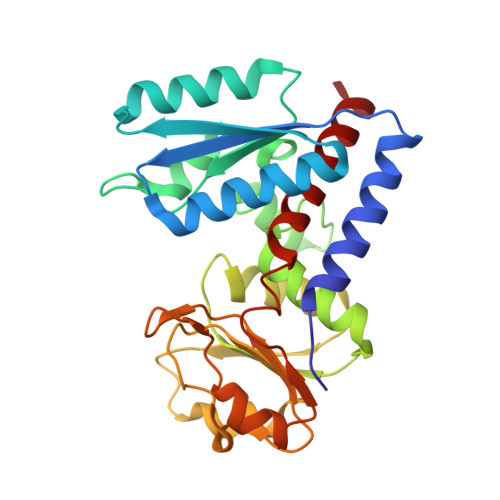
Crystal structure of bifunctional 5,10-methylenetetrahydrofolate dehydrogenase/cyclohydrolase from Thermoplasma acidophilum
Lee, W.H., Sung, M.W., Kim, J.H., Kim, Y.K., Han, A., Hwang, K.Y.(2011) Biochem Biophys Res Commun 406: 459-463
- PubMed: 21333632 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2011.02.074
- Primary Citation Related Structures:
3NGL, 3NGX - PubMed Abstract:
Folate co-enzymes play a pivotal role in one-carbon transfer cellular processes. Many eukaryotes encode the tri-functional tetrahydrofolate dehydrogenase/cyclohydrolase/synthetase (deh/cyc/syn) enzyme, which consists of a N-terminal bifunctional domain (deh/cyc) and a C-terminal monofunctional domain (syn). Here, we report the first analogous archeal enzyme structures, for the bifunctional methylenetetrahydrofolate dehydrogenase/cyclohydrolase from Thermoplasma acidophilum (TaMTHFDC) as the native protein and also as its NADP complex. The TaMTHFDC structure is a dimer with a polar interface, as well as a NADP binding site that shows minor conformational change. The orientations of the residues in the NADP binding site do not change on ligand binding, incorporating three water molecules which are hydrogen bonded with phosphate groups of NADP in the structure of the complex. Our structural information will contribute to an improved understanding of the basis of THF and one-carbon metabolism.
- Division of Biotechnology, College of Life Sciences and Biotechnology, Korea University, Seoul 136-713, South Korea.
Organizational Affiliation: