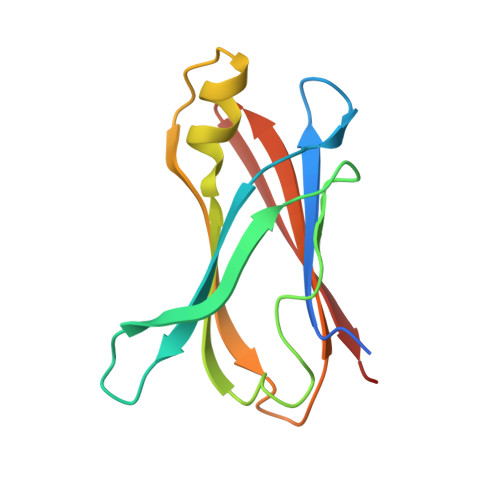
Crystal structure of green tea polyphenol(-)-epigallocatechin gallate (EGCG)-transthyretin complex reveals novel binding site distinct from thyroxine binding site
Miyata, M., Sato, T., Kugimiya, M., Sho, M., Nakamura, T., Ikemizu, S., Chirifu, M., Mizuguchi, M., Nabeshima, Y., Suwa, Y., Morioka, H., Arimori, T., Suico, M.A., Shuto, T., Sako, Y., Momohara, M., Koga, T., Morino-Koga, S., Yamagata, Y., Kai, H.(2010) Biochemistry
- PubMed: 20565072 Search on PubMed
- DOI: https://doi.org/10.1021/bi1004409
- Primary Citation Related Structures:
3NG5 - PubMed Abstract:
Amyloid fibril formation is associated with protein misfolding disorders, including neurodegenerative diseases such as Alzheimer's, Parkinson's, and Huntington's diseases. Familial amyloid polyneuropathy (FAP) is a hereditary disease caused by a point mutation of the human plasma protein, transthyretin (TTR), which binds and transports thyroxine (T(4)). TTR variants contribute to the pathogenesis of amyloidosis by forming amyloid fibrils in the extracellular environment. A recent report showed that epigallocatechin 3-gallate (EGCG), the major polyphenol component of green tea, binds to TTR and suppresses TTR amyloid fibril formation. However, structural analysis of EGCG binding to TTR has not yet been conducted. Here we first investigated the crystal structure of the EGCG-V30M TTR complex and found novel binding sites distinct from the thyroxine binding site, suggesting that EGCG has a mode of action different from those of previous chemical compounds that were shown to bind and stabilize the TTR tetramer structure. Furthermore, EGCG induced the oligomerization and monomer suppression in the cellular system of clinically reported TTR variants. Taken together, these findings suggest the possibility that EGCG may be a candidate compound for FAP therapy.
- Department of Molecular Medicine, Global COE Cell Fate Regulation Research and Education Unit, Graduate School of Pharmaceutical Sciences, Kumamoto University, 5-1 Oe-Honmachi, Kumamoto 862-0973, Japan.
Organizational Affiliation: