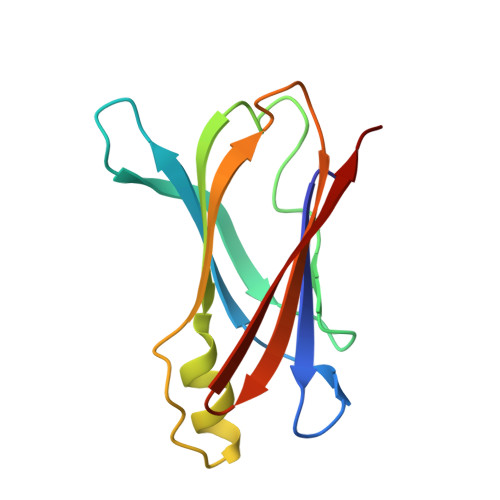
The binding of synthetic triiodo l-thyronine analogs to human transthyretin: molecular basis of cooperative and non-cooperative ligand recognition.
Trivella, D.B., Sairre, M.I., Foguel, D., Lima, L.M., Polikarpov, I.(2011) J Struct Biol 173: 323-332
- PubMed: 20937391
- DOI: https://doi.org/10.1016/j.jsb.2010.10.003
- Primary Citation Related Structures:
3NEE, 3NEO, 3NES, 3NEX - PubMed Abstract:
Transthyretin (TTR) is a tetrameric β-sheet-rich transporter protein directly involved in human amyloid diseases. Several classes of small molecules can bind to TTR delaying its amyloid fibril formation, thus being promising drug candidates to treat TTR amyloidoses. In the present study, we characterized the interactions of the synthetic triiodo L-thyronine analogs and thyroid hormone nuclear receptor TRβ-selective agonists GC-1 and GC-24 with the wild type and V30M variant of human transthyretin (TTR). To achieve this aim, we conducted in vitro TTR acid-mediated aggregation and isothermal titration calorimetry experiments and determined the TTR:GC-1 and TTR:GC-24 crystal structures. Our data indicate that both GC-1 and GC-24 bind to TTR in a non-cooperative manner and are good inhibitors of TTR aggregation, with dissociation constants for both hormone binding sites (HBS) in the low micromolar range. Analysis of the crystal structures of TTRwt:GC-1(24) complexes and their comparison with the TTRwt X-ray structure bound to its natural ligand thyroxine (T4) suggests, at the molecular level, the basis for the cooperative process displayed by T4 and the non-cooperative process provoked by both GC-1 and GC-24 during binding to TTR.
- Instituto de Física de São Carlos-Universidade de São Paulo, São Carlos, SP, Brazil.
Organizational Affiliation: