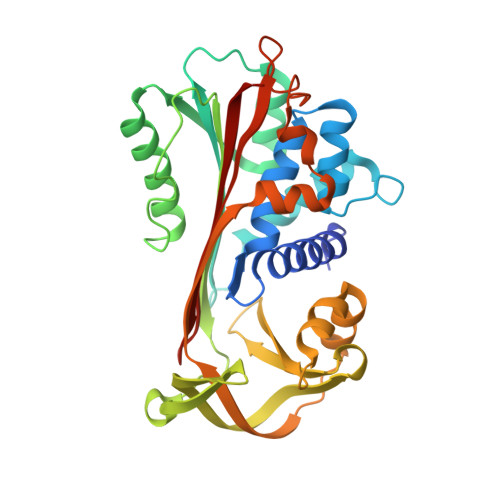
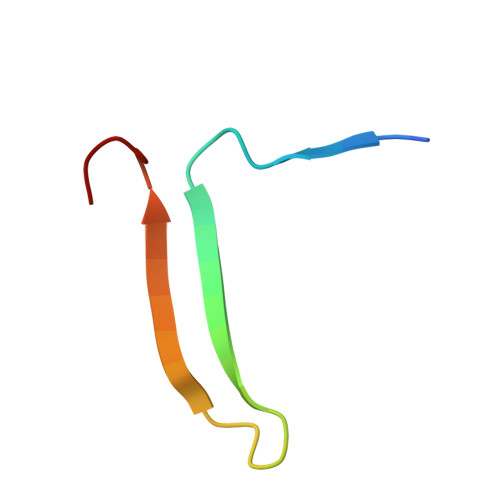
Loop-sheet mechanism of serpin polymerization tested by reactive center loop mutations
Yamasaki, M., Sendall, T.J., Harris, L.E., Lewis, G.M.W., Huntington, J.A.(2010) J Biological Chem 285: 30752-30758
- PubMed: 20667823
- DOI: https://doi.org/10.1074/jbc.M110.156042
- Primary Citation of Related Structures:
3NDD, 3NDF - PubMed Abstract:
The serpin mechanism of protease inhibition involves the rapid and stable incorporation of the reactive center loop (RCL) into central β-sheet A. Serpins therefore require a folding mechanism that bypasses the most stable "loop-inserted" conformation to trap the RCL in an exposed and metastable state. This unusual feature of serpins renders them highly susceptible to point mutations that lead to the accumulation of hyperstable misfolded polymers in the endoplasmic reticulum of secretory cells. The ordered and stable protomer-protomer association in serpin polymers has led to the acceptance of the "loop-sheet" hypothesis of polymerization, where a portion of the RCL of one protomer incorporates in register into sheet A of another. Although this mechanism was proposed 20 years ago, no study has ever been conducted to test its validity. Here, we describe the properties of a variant of α(1)-antitrypsin with a critical hydrophobic section of the RCL substituted with aspartic acid (P8-P6). In contrast to the control, the variant was unable to polymerize when incubated with small peptides or when cleaved in the middle of the RCL (accepted models of loop-sheet polymerization). However, when induced by guanidine HCl or heat, the variant polymerized in a manner indistinguishable from the control. Importantly, the Asp mutations did not affect the ability of the Z or Siiyama α(1)-antitrypsin variants to polymerize in COS-7 cells. These results argue strongly against the loop-sheet hypothesis and suggest that, in serpin polymers, the P8-P6 region is only a small part of an extensive domain swap.
- Department of Haematology, Cambridge Institute for Medical Research, University of Cambridge, Hills Road, Cambridge CB2 0XY, United Kingdom.
Organizational Affiliation: