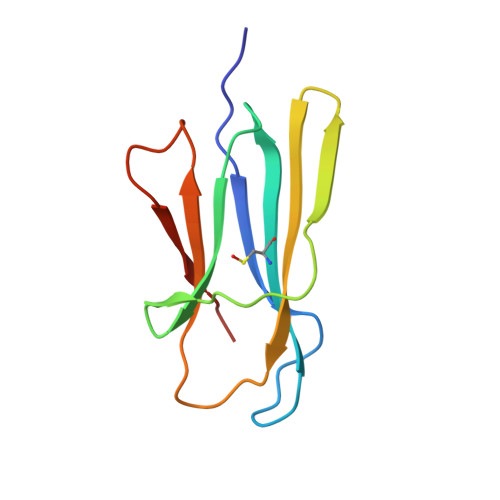
D-strand perturbation and amyloid propensity in beta-2 microglobulin
Azinas, S., Colombo, M., Barbiroli, A., Santambrogio, C., Giorgetti, S., Raimondi, S., Bonomi, F., Grandori, R., Bellotti, V., Ricagno, S., Bolognesi, M.(2011) FEBS J 278: 2349-2358
- PubMed: 21569201 Search on PubMed
- DOI: https://doi.org/10.1111/j.1742-4658.2011.08157.x
- Primary Citation Related Structures:
3NA4 - PubMed Abstract:
Proteins hosting main β-sheets adopt specific strategies to avoid intermolecular interactions leading to aggregation and amyloid deposition. Human beta-2 microglobulin (β2m) displays a typical immunoglobulin fold and is known to be amyloidogenic in vivo. Upon severe kidney deficiency, β2m accumulates in the bloodstream, triggering, over the years, pathological deposition of large amyloid aggregates in joints and bones. A β-bulge observed on the edge D β-strand of some β2m crystal structures has been suggested to be crucial in protecting the protein from amyloid aggregation. Conversely, a straight D-strand, observed in different crystal structures of monomeric β2m, could promote amyloid aggregation. More recently, the different conformations observed for the β2m D-strand have been interpreted as the result of intrinsic flexibility, rather than being assigned to a functional protective role against aggregation. To shed light on such contrasting picture, the mutation Asp53→Pro was engineered in β2m, aiming to impair the formation of a regular/straight D-strand. Such a mutant was characterized structurally and biophysically by CD, X-ray crystallography and MS, in addition to an assessment of its amyloid aggregation trends in vitro. The results reported in the present study highlight the conformational plasticity of the edge D-strand, and show that even perturbing the D-strand structure through a Pro residue has only marginal effects on protecting β2m from amyloid aggregation in vitro.
- Dipartimento di Scienze Biomolecolari e Biotecnologie and CIMAINA, Università degli Studi di Milano, Milan, Italy.
Organizational Affiliation: