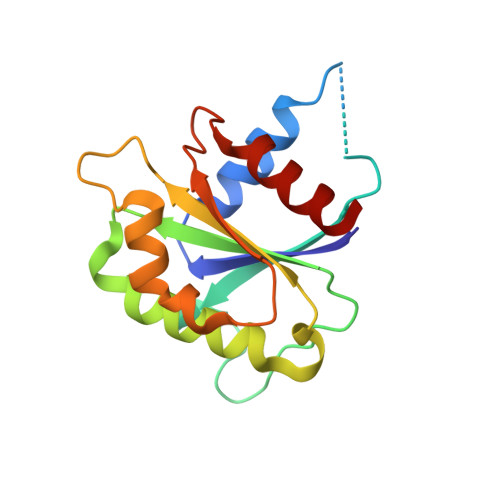
SAXS and X-ray crystallography suggest an unfolding model for the GDP/GTP conformational switch of the small GTPase Arf6.
Biou, V., Aizel, K., Roblin, P., Thureau, A., Jacquet, E., Hansson, S., Guibert, B., Guittet, E., van Heijenoort, C., Zeghouf, M., Perez, J., Cherfils, J.(2010) J Mol Biology 402: 696-707
- PubMed: 20709080 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2010.08.002
- Primary Citation Related Structures:
3N5C - PubMed Abstract:
The small GTPases Arf1 and Arf6 have nonoverlapping functions in cellular traffic despite their very high sequence and structural resemblance. Notably, the exquisite isoform specificity of their guanine nucleotide exchange factors and their distinctive sensitivity to the drug brefeldin A cannot be explained by any straightforward structural model. Here we integrated structural and spectroscopic methods to address this issue using Δ13Arf6-GDP, a truncated mutant that mimics membrane-bound Arf6-GDP. The crystal structure of Δ13Arf6-GDP reveals an unprecedented unfolding of the GTPase core β-strands, which is fully accounted for by small-angle X-ray scattering data in solution and by ab initio three-dimensional envelope calculation. NMR chemical shifts identify this structural disorder in Δ13Arf6-GDP, but not in the closely related Δ17Arf1-GDP, which is consistent with their comparative thermodynamic and hydrodynamic analyses. Taken together, these experiments suggest an unfolding model for the nucleotide switch of Arf6 and shed new light on its biochemical differences with Arf1.
- Laboratoire d'Enzymologie et Biochimie Structurales, Centre de Recherche CNRS de Gif-Sur-Yvette, Gif-sur-Yvette, France. biou@lebs.cnrs-gif.fr
Organizational Affiliation: