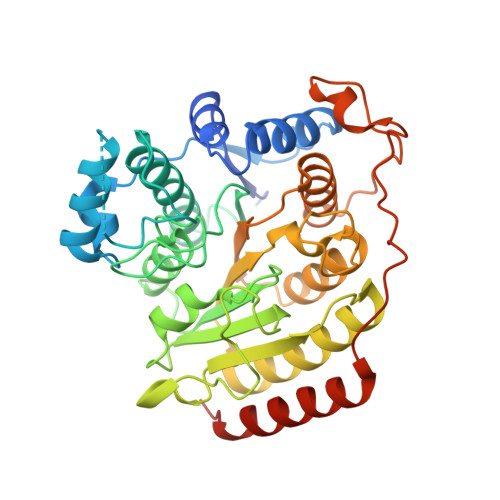
Structures of metal-substituted human histone deacetylase 8 provide mechanistic inferences on biological function.
Dowling, D.P., Gattis, S.G., Fierke, C.A., Christianson, D.W.(2010) Biochemistry 49: 5048-5056
- PubMed: 20545365 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi1005046
- Primary Citation Related Structures:
3MZ3, 3MZ4, 3MZ6, 3MZ7 - PubMed Abstract:
The metal-dependent histone deacetylases (HDACs) adopt an alpha/beta protein fold first identified in rat liver arginase. Despite insignificant overall amino acid sequence identity, these enzymes share a strictly conserved metal binding site with divergent metal specificity and stoichiometry. HDAC8, originally thought to be a Zn(2+)-metallohydrolase, exhibits increased activity with Co(2+) and Fe(2+) cofactors based on k(cat)/K(M) (Gantt, S. L., Gattis, S. G., and Fierke, C. A. (2006) Biochemistry 45, 6170-6178). Here, we report the first X-ray crystal structures of metallo-substituted HDAC8, Co(2+)-HDAC8, D101L Co(2+)-HDAC8, D101L Mn(2+)-HDAC8, and D101L Fe(2+)-HDAC8, each complexed with the inhibitor M344. Metal content of protein samples in solution is confirmed by inductively coupled plasma mass spectrometry. For the crystalline enzymes, peaks in Bijvoet difference Fourier maps calculated from X-ray diffraction data collected near the respective elemental absorption edges confirm metal substitution. Additional solution studies confirm incorporation of Cu(2+); Fe(3+) and Ni(2+) do not bind under conditions tested. The metal dependence of the substrate K(M) values and the K(i) values of hydroxamate inhibitors that chelate the active site metal are consistent with substrate-metal coordination in the precatalytic Michaelis complex that enhances catalysis. Additionally, although HDAC8 binds Zn(2+) nearly 10(6)-fold more tightly than Fe(2+), the affinities for both metal ions are comparable to the readily exchangeable metal concentrations estimated in living cells, suggesting that HDAC8 could bind either or both Fe(2+) or Zn(2+) in vivo.
- Roy and Diana Vagelos Laboratories, Department of Chemistry, University of Pennsylvania, Philadelphia, Pennsylvania 19104-6323, USA.
Organizational Affiliation: