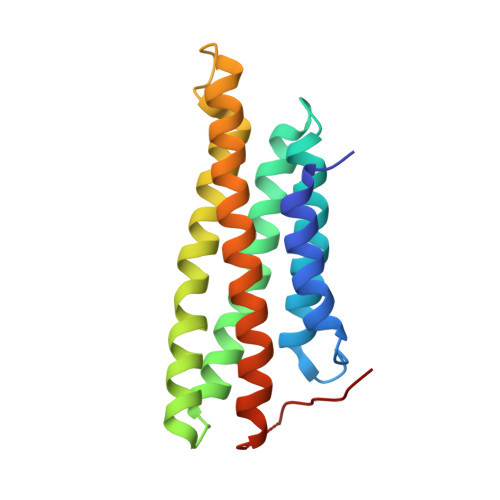
A helix replacement mechanism directs metavinculin functions.
Rangarajan, E.S., Lee, J.H., Yogesha, S.D., Izard, T.(2010) PLoS One 5: e10679-e10679
- PubMed: 20502710 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0010679
- Primary Citation Related Structures:
3MYI - PubMed Abstract:
Cells require distinct adhesion complexes to form contacts with their neighbors or the extracellular matrix, and vinculin links these complexes to the actin cytoskeleton. Metavinculin, an isoform of vinculin that harbors a unique 68-residue insert in its tail domain, has distinct actin bundling and oligomerization properties and plays essential roles in muscle development and homeostasis. Moreover, patients with sporadic or familial mutations in the metavinculin-specific insert invariably develop fatal cardiomyopathies. Here we report the high resolution crystal structure of the metavinculin tail domain, as well as the crystal structures of full-length human native metavinculin (1,134 residues) and of the full-length cardiomyopathy-associated DeltaLeu954 metavinculin deletion mutant. These structures reveal that an alpha-helix (H1') and extended coil of the metavinculin insert replace alpha-helix H1 and its preceding extended coil found in the N-terminal region of the vinculin tail domain to form a new five-helix bundle tail domain. Further, biochemical analyses demonstrate that this helix replacement directs the distinct actin bundling and oligomerization properties of metavinculin. Finally, the cardiomyopathy associated DeltaLeu954 and Arg975Trp metavinculin mutants reside on the replaced extended coil and the H1' alpha-helix, respectively. Thus, a helix replacement mechanism directs metavinculin's unique functions.
- Cell Adhesion Laboratory, Department of Cancer Biology, The Scripps Research Institute, Scripps Florida, Jupiter, Florida, United States of America.
Organizational Affiliation: