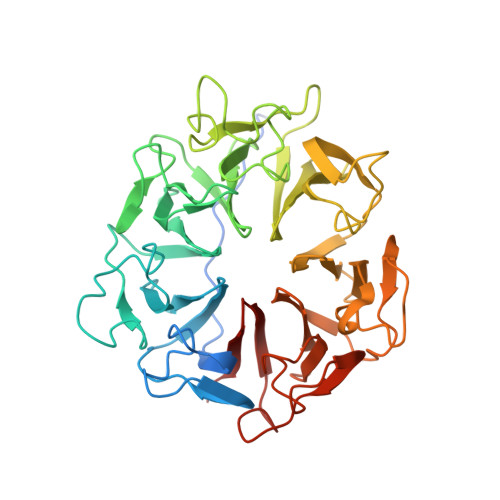
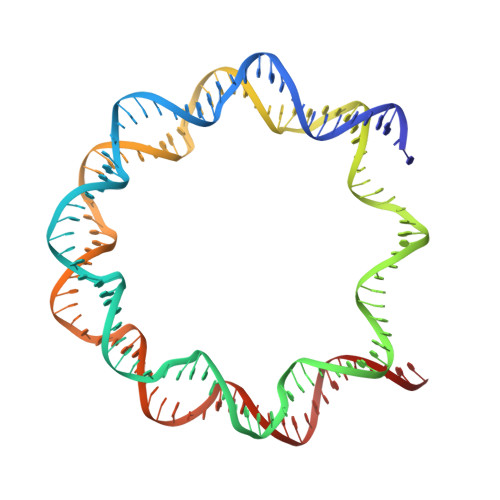
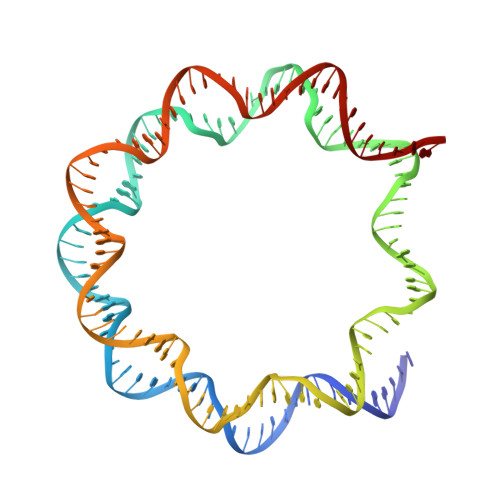
Structure of RCC1 chromatin factor bound to the nucleosome core particle.
Makde, R.D., England, J.R., Yennawar, H.P., Tan, S.(2010) Nature 467: 562-566
- PubMed: 20739938 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature09321
- Primary Citation Related Structures:
3MVD - PubMed Abstract:
The small GTPase Ran enzyme regulates critical eukaryotic cellular functions including nuclear transport and mitosis through the creation of a RanGTP gradient around the chromosomes. This concentration gradient is created by the chromatin-bound RCC1 (regulator of chromosome condensation) protein, which recruits Ran to nucleosomes and activates Ran's nucleotide exchange activity. Although RCC1 has been shown to bind directly with the nucleosome, the molecular details of this interaction were not known. Here we determine the crystal structure of a complex of Drosophila RCC1 and the nucleosome core particle at 2.9 Å resolution, providing an atomic view of how a chromatin protein interacts with the histone and DNA components of the nucleosome. Our structure also suggests that the Widom 601 DNA positioning sequence present in the nucleosomes forms a 145-base-pair nucleosome core particle, not the expected canonical 147-base-pair particle.
- Center for Eukaryotic Gene Regulation, Department of Biochemistry & Molecular Biology, The Pennsylvania State University, University Park, Pennsylvania 16802, USA.
Organizational Affiliation: