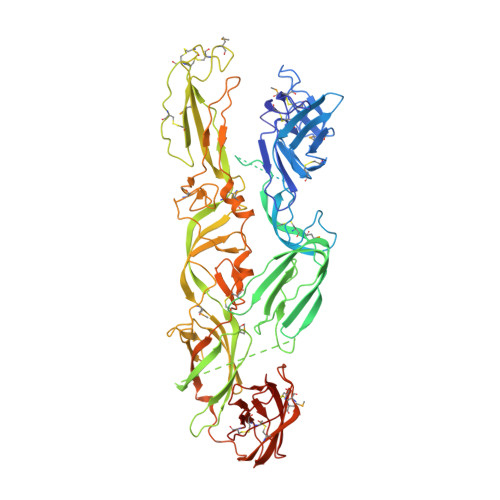
Structural changes of envelope proteins during alphavirus fusion.
Li, L., Jose, J., Xiang, Y., Kuhn, R.J., Rossmann, M.G.(2010) Nature 468: 705-708
- PubMed: 21124457 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature09546
- Primary Citation Related Structures:
3MUU, 3MUW - PubMed Abstract:
Alphaviruses are enveloped RNA viruses that have a diameter of about 700 Å and can be lethal human pathogens. Entry of virus into host cells by endocytosis is controlled by two envelope glycoproteins, E1 and E2. The E2-E1 heterodimers form 80 trimeric spikes on the icosahedral virus surface, 60 with quasi-three-fold symmetry and 20 coincident with the icosahedral three-fold axes arranged with T = 4 quasi-symmetry. The E1 glycoprotein has a hydrophobic fusion loop at one end and is responsible for membrane fusion. The E2 protein is responsible for receptor binding and protects the fusion loop at neutral pH. The lower pH in the endosome induces the virions to undergo an irreversible conformational change in which E2 and E1 dissociate and E1 forms homotrimers, triggering fusion of the viral membrane with the endosomal membrane and then releasing the viral genome into the cytoplasm. Here we report the structure of an alphavirus spike, crystallized at low pH, representing an intermediate in the fusion process and clarifying the maturation process. The trimer of E2-E1 in the crystal structure is similar to the spikes in the neutral pH virus except that the E2 middle region is disordered, exposing the fusion loop. The amino- and carboxy-terminal domains of E2 each form immunoglobulin-like folds, consistent with the receptor attachment properties of E2.
- Department of Biological Sciences, Purdue University, 915 W. State Street, West Lafayette, Indiana 47907-2054, USA.
Organizational Affiliation: